Nazim Bouatta
QuickBind: A Light-Weight And Interpretable Molecular Docking Model
Oct 21, 2024
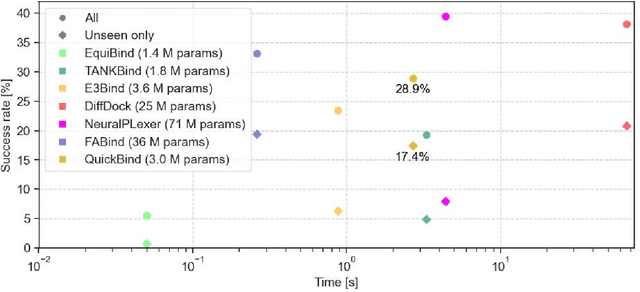
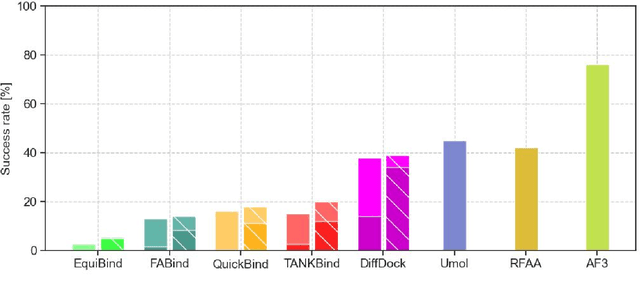

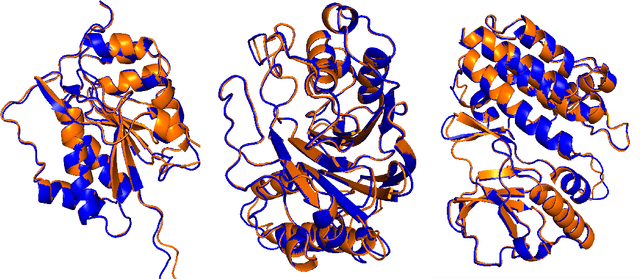
Abstract:Predicting a ligand's bound pose to a target protein is a key component of early-stage computational drug discovery. Recent developments in machine learning methods have focused on improving pose quality at the cost of model runtime. For high-throughput virtual screening applications, this exposes a capability gap that can be filled by moderately accurate but fast pose prediction. To this end, we developed QuickBind, a light-weight pose prediction algorithm. We assess QuickBind on widely used benchmarks and find that it provides an attractive trade-off between model accuracy and runtime. To facilitate virtual screening applications, we augment QuickBind with a binding affinity module and demonstrate its capabilities for multiple clinically-relevant drug targets. Finally, we investigate the mechanistic basis by which QuickBind makes predictions and find that it has learned key physicochemical properties of molecular docking, providing new insights into how machine learning models generate protein-ligand poses. By virtue of its simplicity, QuickBind can serve as both an effective virtual screening tool and a minimal test bed for exploring new model architectures and innovations. Model code and weights are available at https://github.com/aqlaboratory/QuickBind .
OpenProteinSet: Training data for structural biology at scale
Aug 10, 2023


Abstract:Multiple sequence alignments (MSAs) of proteins encode rich biological information and have been workhorses in bioinformatic methods for tasks like protein design and protein structure prediction for decades. Recent breakthroughs like AlphaFold2 that use transformers to attend directly over large quantities of raw MSAs have reaffirmed their importance. Generation of MSAs is highly computationally intensive, however, and no datasets comparable to those used to train AlphaFold2 have been made available to the research community, hindering progress in machine learning for proteins. To remedy this problem, we introduce OpenProteinSet, an open-source corpus of more than 16 million MSAs, associated structural homologs from the Protein Data Bank, and AlphaFold2 protein structure predictions. We have previously demonstrated the utility of OpenProteinSet by successfully retraining AlphaFold2 on it. We expect OpenProteinSet to be broadly useful as training and validation data for 1) diverse tasks focused on protein structure, function, and design and 2) large-scale multimodal machine learning research.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge