Nathaniel Braman
Novel Radiomic Measurements of Tumor- Associated Vasculature Morphology on Clinical Imaging as a Biomarker of Treatment Response in Multiple Cancers
Oct 05, 2022Abstract:Purpose: Tumor-associated vasculature differs from healthy blood vessels by its chaotic architecture and twistedness, which promotes treatment resistance. Measurable differences in these attributes may help stratify patients by likely benefit of systemic therapy (e.g. chemotherapy). In this work, we present a new category of radiomic biomarkers called quantitative tumor-associated vasculature (QuanTAV) features, and demonstrate their ability to predict response and survival across multiple cancers, imaging modalities, and treatment regimens. Experimental Design: We segmented tumor vessels and computed mathematical measurements of twistedness and organization on routine pre-treatment radiology (CT or contrast-enhanced MRI) from 558 patients, who received one of four first-line chemotherapy-based therapeutic intervention strategies for breast (n=371) or non-small cell lung cancer (NSCLC, n=187). Results: Across 4 chemotherapy-based treatment strategies, classifiers of QuanTAV measurements significantly (p<.05) predicted response in held out testing cohorts alone (AUC=0.63-0.71) and increased AUC by 0.06-0.12 when added to models of significant clinical variables alone. QuanTAV risk scores were prognostic of recurrence free survival in treatment cohorts chemotherapy for breast cancer (p=0.002, HR=1.25, 95% CI 1.08-1.44, C-index=.66) and chemoradiation for NSCLC (p=0.039, HR=1.28, 95% CI 1.01-1.62, C-index=0.66). Categorical QuanTAV risk groups were independently prognostic among all treatment groups, including NSCLC patients receiving chemotherapy (p=0.034, HR=2.29, 95% CI 1.07-4.94, C-index=0.62). Conclusions: Across these domains, we observed an association of vascular morphology on radiology with treatment outcome. Our findings suggest the potential of tumor-associated vasculature shape and structure as a prognostic and predictive biomarker for multiple cancers and treatments.
Deep Orthogonal Fusion: Multimodal Prognostic Biomarker Discovery Integrating Radiology, Pathology, Genomic, and Clinical Data
Jul 01, 2021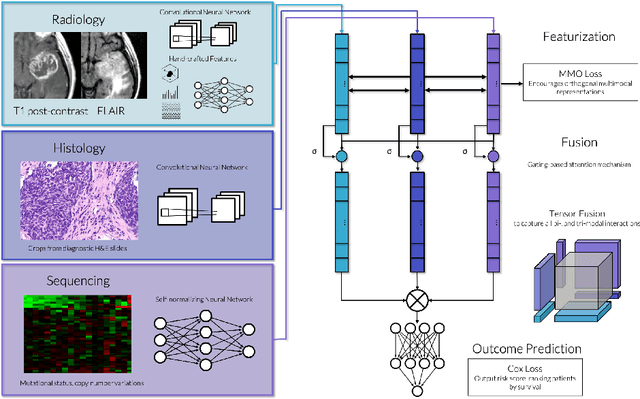
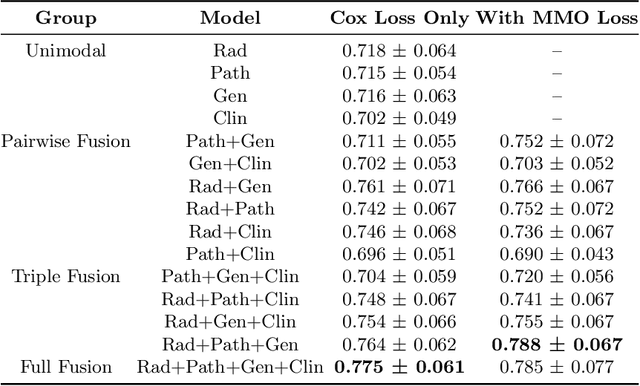
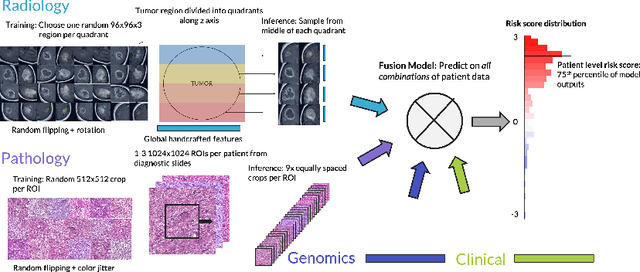
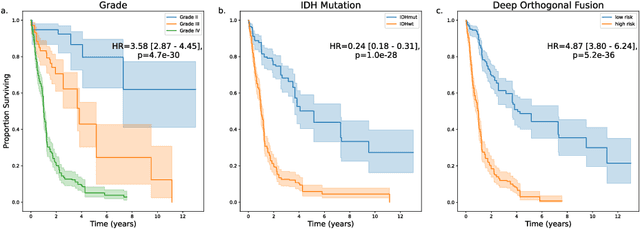
Abstract:Clinical decision-making in oncology involves multimodal data such as radiology scans, molecular profiling, histopathology slides, and clinical factors. Despite the importance of these modalities individually, no deep learning framework to date has combined them all to predict patient prognosis. Here, we predict the overall survival (OS) of glioma patients from diverse multimodal data with a Deep Orthogonal Fusion (DOF) model. The model learns to combine information from multiparametric MRI exams, biopsy-based modalities (such as H&E slide images and/or DNA sequencing), and clinical variables into a comprehensive multimodal risk score. Prognostic embeddings from each modality are learned and combined via attention-gated tensor fusion. To maximize the information gleaned from each modality, we introduce a multimodal orthogonalization (MMO) loss term that increases model performance by incentivizing constituent embeddings to be more complementary. DOF predicts OS in glioma patients with a median C-index of 0.788 +/- 0.067, significantly outperforming (p=0.023) the best performing unimodal model with a median C-index of 0.718 +/- 0.064. The prognostic model significantly stratifies glioma patients by OS within clinical subsets, adding further granularity to prognostic clinical grading and molecular subtyping.
Deep learning-based prediction of response to HER2-targeted neoadjuvant chemotherapy from pre-treatment dynamic breast MRI: A multi-institutional validation study
Jan 22, 2020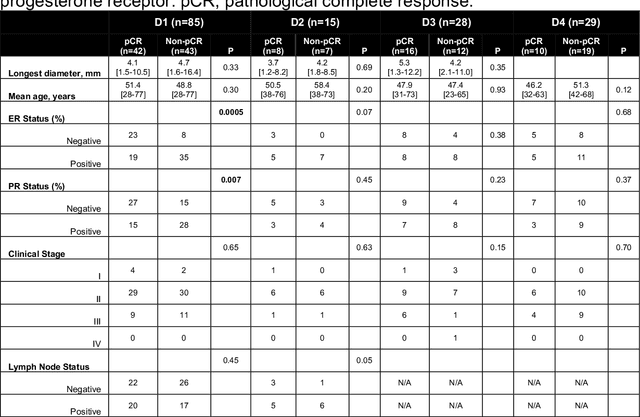
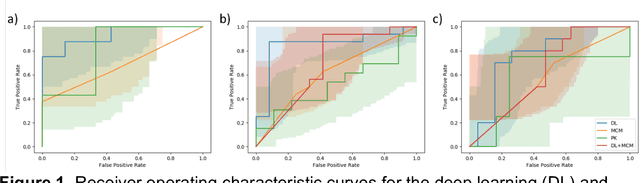
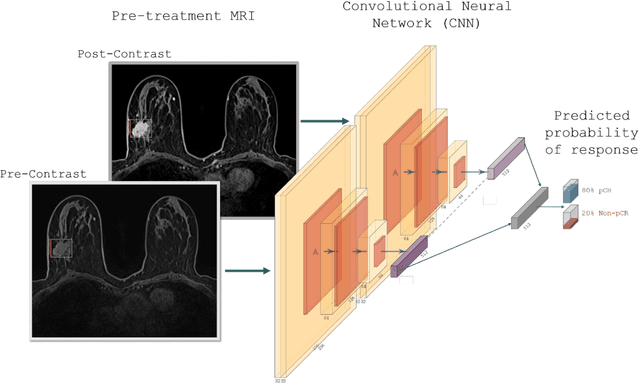
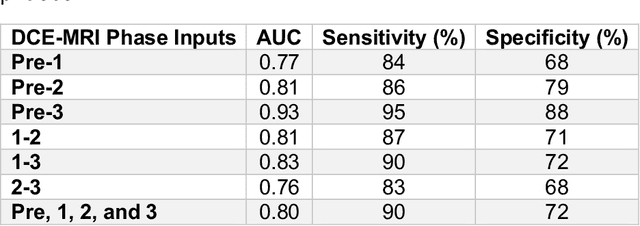
Abstract:Predicting response to neoadjuvant therapy is a vexing challenge in breast cancer. In this study, we evaluate the ability of deep learning to predict response to HER2-targeted neo-adjuvant chemotherapy (NAC) from pre-treatment dynamic contrast-enhanced (DCE) MRI acquired prior to treatment. In a retrospective study encompassing DCE-MRI data from a total of 157 HER2+ breast cancer patients from 5 institutions, we developed and validated a deep learning approach for predicting pathological complete response (pCR) to HER2-targeted NAC prior to treatment. 100 patients who received HER2-targeted neoadjuvant chemotherapy at a single institution were used to train (n=85) and tune (n=15) a convolutional neural network (CNN) to predict pCR. A multi-input CNN leveraging both pre-contrast and late post-contrast DCE-MRI acquisitions was identified to achieve optimal response prediction within the validation set (AUC=0.93). This model was then tested on two independent testing cohorts with pre-treatment DCE-MRI data. It achieved strong performance in a 28 patient testing set from a second institution (AUC=0.85, 95% CI 0.67-1.0, p=.0008) and a 29 patient multicenter trial including data from 3 additional institutions (AUC=0.77, 95% CI 0.58-0.97, p=0.006). Deep learning-based response prediction model was found to exceed a multivariable model incorporating predictive clinical variables (AUC < .65 in testing cohorts) and a model of semi-quantitative DCE-MRI pharmacokinetic measurements (AUC < .60 in testing cohorts). The results presented in this work across multiple sites suggest that with further validation deep learning could provide an effective and reliable tool to guide targeted therapy in breast cancer, thus reducing overtreatment among HER2+ patients.
Disease Detection in Weakly Annotated Volumetric Medical Images using a Convolutional LSTM Network
Dec 03, 2018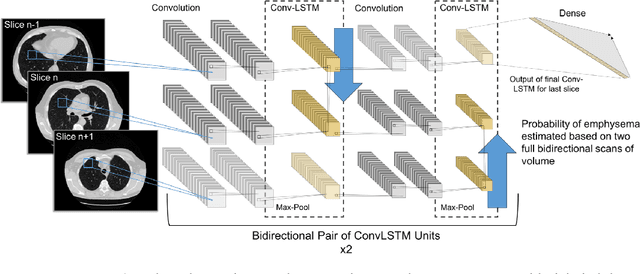
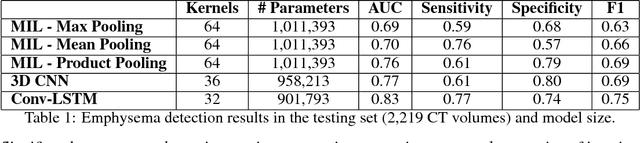
Abstract:We explore a solution for learning disease signatures from weakly, yet easily obtainable, annotated volumetric medical imaging data by analyzing 3D volumes as a sequence of 2D images. We demonstrate the performance of our solution in the detection of emphysema in lung cancer screening low-dose CT images. Our approach utilizes convolutional long short-term memory (LSTM) to "scan" sequentially through an imaging volume for the presence of disease in a portion of scanned region. This framework allowed effective learning given only volumetric images and binary disease labels, thus enabling training from a large dataset of 6,631 un-annotated image volumes from 4,486 patients. When evaluated in a testing set of 2,163 volumes from 2,163 patients, our model distinguished emphysema with area under the receiver operating characteristic curve (AUC) of .83. This approach was found to outperform 2D convolutional neural networks (CNN) implemented with various multiple-instance learning schemes (AUC=0.69-0.76) and a 3D CNN (AUC=.77).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge