Mohammed Amine Gharsallaoui
Streamflow Prediction with Uncertainty Quantification for Water Management: A Constrained Reasoning and Learning Approach
May 31, 2024Abstract:Predicting the spatiotemporal variation in streamflow along with uncertainty quantification enables decision-making for sustainable management of scarce water resources. Process-based hydrological models (aka physics-based models) are based on physical laws, but using simplifying assumptions which can lead to poor accuracy. Data-driven approaches offer a powerful alternative, but they require large amount of training data and tend to produce predictions that are inconsistent with physical laws. This paper studies a constrained reasoning and learning (CRL) approach where physical laws represented as logical constraints are integrated as a layer in the deep neural network. To address small data setting, we develop a theoretically-grounded training approach to improve the generalization accuracy of deep models. For uncertainty quantification, we combine the synergistic strengths of Gaussian processes (GPs) and deep temporal models (i.e., deep models for time-series forecasting) by passing the learned latent representation as input to a standard distance-based kernel. Experiments on multiple real-world datasets demonstrate the effectiveness of both CRL and GP with deep kernel approaches over strong baseline methods.
Quantifying the Reproducibility of Graph Neural Networks using Multigraph Brain Data
Sep 06, 2021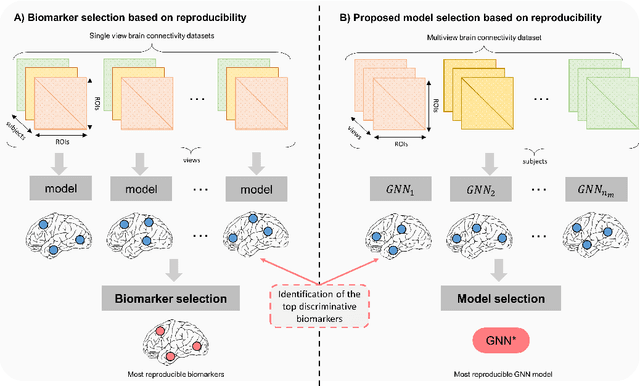



Abstract:Graph neural networks (GNNs) have witnessed an unprecedented proliferation in tackling several problems in computer vision, computer-aided diagnosis, and related fields. While prior studies have focused on boosting the model accuracy, quantifying the reproducibility of the most discriminative features identified by GNNs is still an intact problem that yields concerns about their reliability in clinical applications in particular. Specifically, the reproducibility of biological markers across clinical datasets and distribution shifts across classes (e.g., healthy and disordered brains) is of paramount importance in revealing the underpinning mechanisms of diseases as well as propelling the development of personalized treatment. Motivated by these issues, we propose, for the first time, reproducibility-based GNN selection (RG-Select), a framework for GNN reproducibility assessment via the quantification of the most discriminative features (i.e., biomarkers) shared between different models. To ascertain the soundness of our framework, the reproducibility assessment embraces variations of different factors such as training strategies and data perturbations. Despite these challenges, our framework successfully yielded replicable conclusions across different training strategies and various clinical datasets. Our findings could thus pave the way for the development of biomarker trustworthiness and reliability assessment methods for computer-aided diagnosis and prognosis tasks. RG-Select code is available on GitHub at https://github.com/basiralab/RG-Select.
Predicting cognitive scores with graph neural networks through sample selection learning
Jun 17, 2021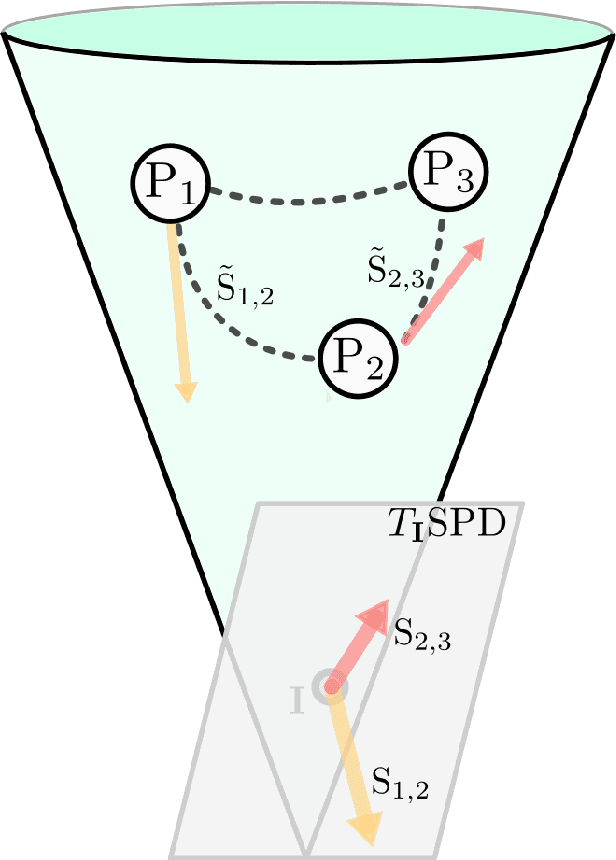
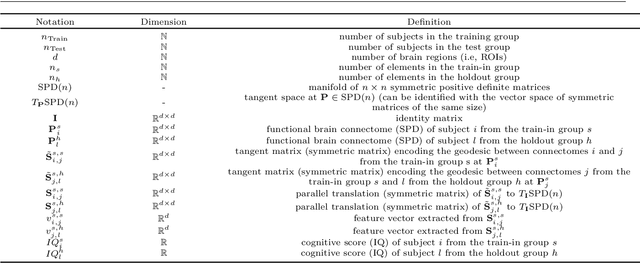
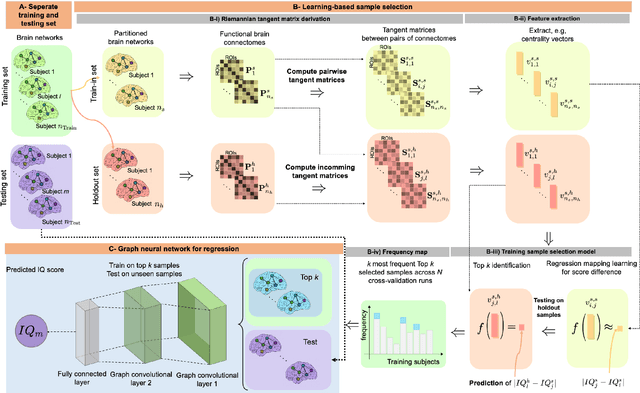
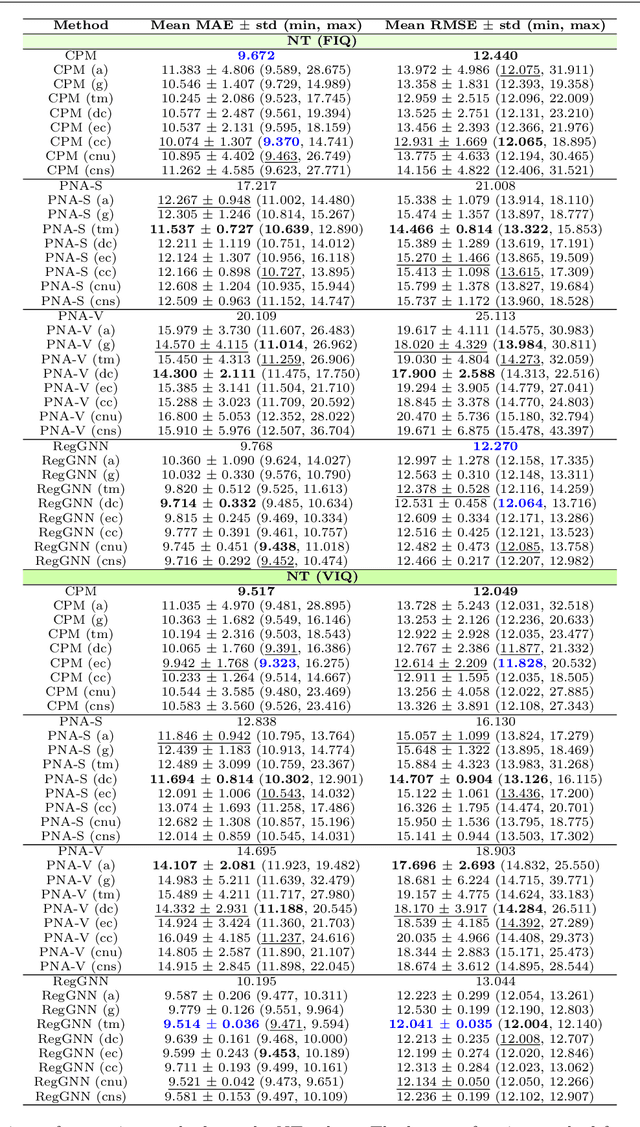
Abstract:Analyzing the relation between intelligence and neural activity is of the utmost importance in understanding the working principles of the human brain in health and disease. In existing literature, functional brain connectomes have been used successfully to predict cognitive measures such as intelligence quotient (IQ) scores in both healthy and disordered cohorts using machine learning models. However, existing methods resort to flattening the brain connectome (i.e., graph) through vectorization which overlooks its topological properties. To address this limitation and inspired from the emerging graph neural networks (GNNs), we design a novel regression GNN model (namely RegGNN) for predicting IQ scores from brain connectivity. On top of that, we introduce a novel, fully modular sample selection method to select the best samples to learn from for our target prediction task. However, since such deep learning architectures are computationally expensive to train, we further propose a \emph{learning-based sample selection} method that learns how to choose the training samples with the highest expected predictive power on unseen samples. For this, we capitalize on the fact that connectomes (i.e., their adjacency matrices) lie in the symmetric positive definite (SPD) matrix cone. Our results on full-scale and verbal IQ prediction outperforms comparison methods in autism spectrum disorder cohorts and achieves a competitive performance for neurotypical subjects using 3-fold cross-validation. Furthermore, we show that our sample selection approach generalizes to other learning-based methods, which shows its usefulness beyond our GNN architecture.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge