Lovro Vrček
Learning to Untangle Genome Assembly with Graph Convolutional Networks
Jun 01, 2022
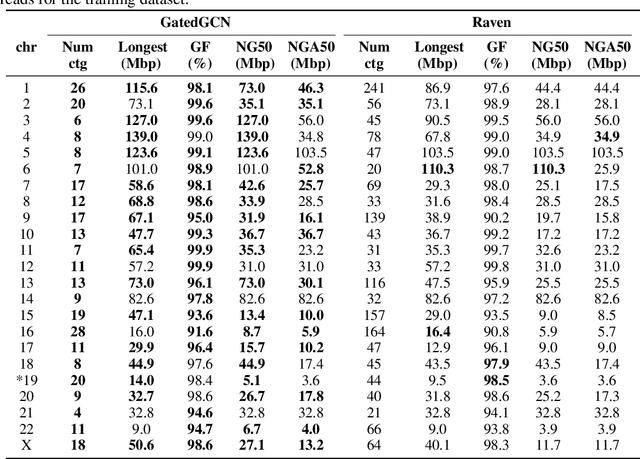
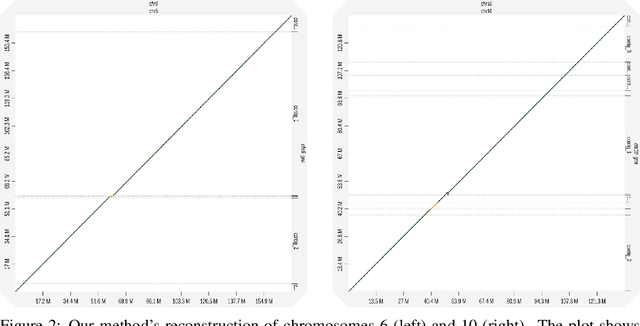
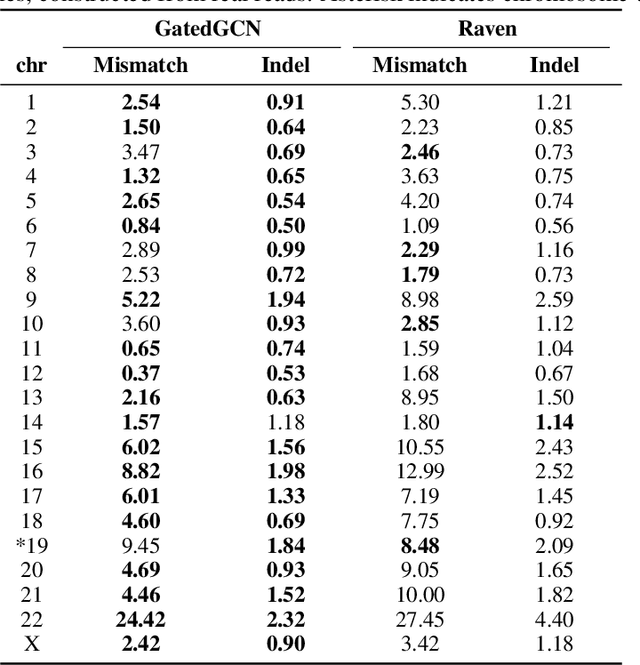
Abstract:A quest to determine the complete sequence of a human DNA from telomere to telomere started three decades ago and was finally completed in 2021. This accomplishment was a result of a tremendous effort of numerous experts who engineered various tools and performed laborious manual inspection to achieve the first gapless genome sequence. However, such method can hardly be used as a general approach to assemble different genomes, especially when the assembly speed is critical given the large amount of data. In this work, we explore a different approach to the central part of the genome assembly task that consists of untangling a large assembly graph from which a genomic sequence needs to be reconstructed. Our main motivation is to reduce human-engineered heuristics and use deep learning to develop more generalizable reconstruction techniques. Precisely, we introduce a new learning framework to train a graph convolutional network to resolve assembly graphs by finding a correct path through them. The training is supervised with a dataset generated from the resolved CHM13 human sequence and tested on assembly graphs built using real human PacBio HiFi reads. Experimental results show that a model, trained on simulated graphs generated solely from a single chromosome, is able to remarkably resolve all other chromosomes. Moreover, the model outperforms hand-crafted heuristics from a state-of-the-art \textit{de novo} assembler on the same graphs. Reconstructed chromosomes with graph networks are more accurate on nucleotide level, report lower number of contigs, higher genome reconstructed fraction and NG50/NGA50 assessment metrics.
A step towards neural genome assembly
Nov 10, 2020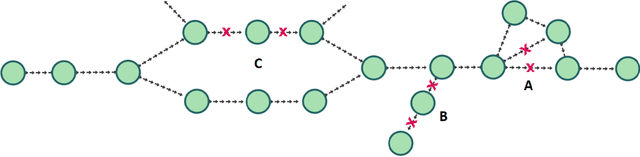
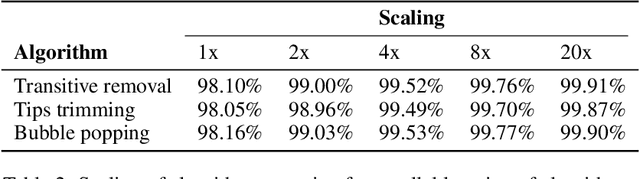
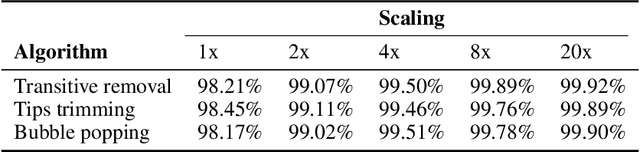

Abstract:De novo genome assembly focuses on finding connections between a vast amount of short sequences in order to reconstruct the original genome. The central problem of genome assembly could be described as finding a Hamiltonian path through a large directed graph with a constraint that an unknown number of nodes and edges should be avoided. However, due to local structures in the graph and biological features, the problem can be reduced to graph simplification, which includes removal of redundant information. Motivated by recent advancements in graph representation learning and neural execution of algorithms, in this work we train the MPNN model with max-aggregator to execute several algorithms for graph simplification. We show that the algorithms were learned successfully and can be scaled to graphs of sizes up to 20 times larger than the ones used in training. We also test on graphs obtained from real-world genomic data---that of a lambda phage and E. coli.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge