Liesbeth Hondelink
Predicting Targeted Therapy Resistance in Non-Small Cell Lung Cancer Using Multimodal Machine Learning
Mar 31, 2025Abstract:Lung cancer is the primary cause of cancer death globally, with non-small cell lung cancer (NSCLC) emerging as its most prevalent subtype. Among NSCLC patients, approximately 32.3% have mutations in the epidermal growth factor receptor (EGFR) gene. Osimertinib, a third-generation EGFR-tyrosine kinase inhibitor (TKI), has demonstrated remarkable efficacy in the treatment of NSCLC patients with activating and T790M resistance EGFR mutations. Despite its established efficacy, drug resistance poses a significant challenge for patients to fully benefit from osimertinib. The absence of a standard tool to accurately predict TKI resistance, including that of osimertinib, remains a critical obstacle. To bridge this gap, in this study, we developed an interpretable multimodal machine learning model designed to predict patient resistance to osimertinib among late-stage NSCLC patients with activating EGFR mutations, achieving a c-index of 0.82 on a multi-institutional dataset. This machine learning model harnesses readily available data routinely collected during patient visits and medical assessments to facilitate precision lung cancer management and informed treatment decisions. By integrating various data types such as histology images, next generation sequencing (NGS) data, demographics data, and clinical records, our multimodal model can generate well-informed recommendations. Our experiment results also demonstrated the superior performance of the multimodal model over single modality models (c-index 0.82 compared with 0.75 and 0.77), thus underscoring the benefit of combining multiple modalities in patient outcome prediction.
Vision Transformer-Based Deep Learning for Histologic Classification of Endometrial Cancer
Dec 13, 2023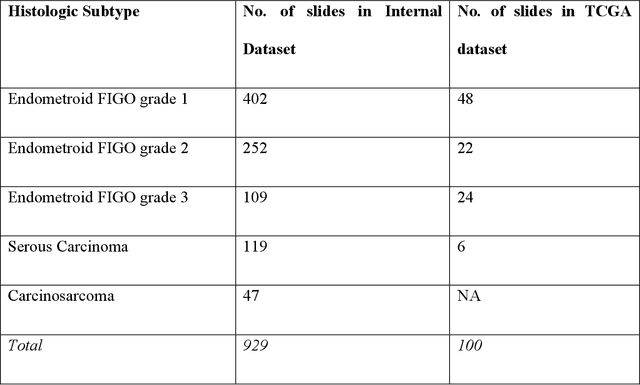



Abstract:Endometrial cancer, the sixth most common cancer in females worldwide, presents as a heterogeneous group with certain types prone to recurrence. Precise histologic evaluation of endometrial cancer is essential for effective patient management and determining the best treatment modalities. This study introduces EndoNet, a transformer-based deep learning approach for histologic classification of endometrial cancer. EndoNet uses convolutional neural networks for extracting histologic features and a vision transformer for aggregating these features and classifying slides based on their visual characteristics. The model was trained on 929 digitized hematoxylin and eosin-stained whole slide images of endometrial cancer from hysterectomy cases at Dartmouth Health. It classifies these slides into low grade (Endometroid Grades 1 and 2) and high-grade (endometroid carcinoma FIGO grade 3, uterine serous carcinoma, carcinosarcoma) categories. EndoNet was evaluated on an internal test set of 218 slides and an external test set of 100 random slides from the public TCGA database. The model achieved a weighted average F1-score of 0.92 (95% CI: 0.87-0.95) and an AUC of 0.93 (95% CI: 0.88-0.96) on the internal test, and 0.86 (95% CI: 0.80-0.94) for F1-score and 0.86 (95% CI: 0.75-0.93) for AUC on the external test. Pending further validation, EndoNet has the potential to assist pathologists in classifying challenging gynecologic pathology tumors and enhancing patient care.
Masked Pre-Training of Transformers for Histology Image Analysis
Apr 14, 2023Abstract:In digital pathology, whole slide images (WSIs) are widely used for applications such as cancer diagnosis and prognosis prediction. Visual transformer models have recently emerged as a promising method for encoding large regions of WSIs while preserving spatial relationships among patches. However, due to the large number of model parameters and limited labeled data, applying transformer models to WSIs remains challenging. Inspired by masked language models, we propose a pretext task for training the transformer model without labeled data to address this problem. Our model, MaskHIT, uses the transformer output to reconstruct masked patches and learn representative histological features based on their positions and visual features. The experimental results demonstrate that MaskHIT surpasses various multiple instance learning approaches by 3% and 2% on survival prediction and cancer subtype classification tasks, respectively. Furthermore, MaskHIT also outperforms two of the most recent state-of-the-art transformer-based methods. Finally, a comparison between the attention maps generated by the MaskHIT model with pathologist's annotations indicates that the model can accurately identify clinically relevant histological structures in each task.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge