Kiwan Jeon
MGMAR: Metal-Guided Metal Artifact Reduction for X-ray Computed Tomography
Mar 13, 2026Abstract:An X-ray computed tomography (CT), metal artifact reduction (MAR) remains a major challenge because metallic implants violate standard CT forward-model assumptions, producing severe streaking and shadowing artifacts that degrade diagnostic quality. We propose MGMAR, a metal-guided MAR method that explicitly leverages metal-related information throughout the reconstruction pipeline. MGMAR first generates a high-quality prior image by training a conditioned implicit neural representation (INR) using metal-unaffected projections, and then incorporates this prior into a normalized MAR (NMAR) framework for projection completion. To improve robustness under severe metal corruption, we pretrain the encoder-conditioned INR on paired metal-corrupted and artifact-free CT images, thereby embedding data-driven prior knowledge into the INR parameter space. This prior-embedded initialization reduces sensitivity to random initialization and accelerates convergence during measurement-specific refinement. The encoder takes a metal-corrupted reconstruction together with a recursively constructed metal artifact image, enabling the latent field to capture metal-dependent global artifact patterns. After projection completion using the INR prior, we further suppress residual artifacts using a metal-conditioned correction network, where the metal mask modulates intermediate features via adaptive instance normalization to target metal-dependent secondary artifacts while preserving anatomical structures. Experiments on the public AAPM-MAR benchmark demonstrate that MGMAR achieves state-of-the-art performance, attaining an average final score of 0.89 on 29 clinical test cases.
ReCo-Diff: Residual-Conditioned Deterministic Sampling for Cold Diffusion in Sparse-View CT
Mar 03, 2026Abstract:Cold and generalized diffusion models have recently shown strong potential for sparse-view CT reconstruction by explicitly modeling deterministic degradation processes. However, existing sampling strategies often rely on ad hoc sampling controls or fixed schedules, which remain sensitive to error accumulation and sampling instability. We propose ReCo-Diff, a residual-conditioned diffusion framework that leverages observation residuals through residual-conditioned self-guided sampling. At each sampling step, ReCo-Diff first produces a null (unconditioned) baseline reconstruction and then conditions subsequent predictions on the observation residual between the predicted image and the measured sparse-view input. This residual-driven guidance provides continuous, measurement-aware correction while preserving a deterministic sampling schedule, without requiring heuristic interventions. Experimental results demonstrate that ReCo-Diff consistently outperforms existing cold diffusion sampling baselines, achieving higher reconstruction accuracy, improved stability, and enhanced robustness under severe sparsity.
An Iterative Reconstruction Method for Dental Cone-Beam Computed Tomography with a Truncated Field of View
Aug 11, 2025Abstract:In dental cone-beam computed tomography (CBCT), compact and cost-effective system designs often use small detectors, resulting in a truncated field of view (FOV) that does not fully encompass the patient's head. In iterative reconstruction approaches, the discrepancy between the actual projection and the forward projection within the truncated FOV accumulates over iterations, leading to significant degradation in the reconstructed image quality. In this study, we propose a two-stage approach to mitigate truncation artifacts in dental CBCT. In the first stage, we employ Implicit Neural Representation (INR), leveraging its superior representation power, to generate a prior image over an extended region so that its forward projection fully covers the patient's head. To reduce computational and memory burdens, INR reconstruction is performed with a coarse voxel size. The forward projection of this prior image is then used to estimate the discrepancy due to truncated FOV in the measured projection data. In the second stage, the discrepancy-corrected projection data is utilized in a conventional iterative reconstruction process within the truncated region. Our numerical results demonstrate that the proposed two-grid approach effectively suppresses truncation artifacts, leading to improved CBCT image quality.
Adaptive Multi-resolution Hash-Encoding Framework for INR-based Dental CBCT Reconstruction with Truncated FOV
Jun 14, 2025Abstract:Implicit neural representation (INR), particularly in combination with hash encoding, has recently emerged as a promising approach for computed tomography (CT) image reconstruction. However, directly applying INR techniques to 3D dental cone-beam CT (CBCT) with a truncated field of view (FOV) is challenging. During the training process, if the FOV does not fully encompass the patient's head, a discrepancy arises between the measured projections and the forward projections computed within the truncated domain. This mismatch leads the network to estimate attenuation values inaccurately, producing severe artifacts in the reconstructed images. In this study, we propose a computationally efficient INR-based reconstruction framework that leverages multi-resolution hash encoding for 3D dental CBCT with a truncated FOV. To mitigate truncation artifacts, we train the network over an expanded reconstruction domain that fully encompasses the patient's head. For computational efficiency, we adopt an adaptive training strategy that uses a multi-resolution grid: finer resolution levels and denser sampling inside the truncated FOV, and coarser resolution levels with sparser sampling outside. To maintain consistent input dimensionality of the network across spatially varying resolutions, we introduce an adaptive hash encoder that selectively activates the lower-level features of the hash hierarchy for points outside the truncated FOV. The proposed method with an extended FOV effectively mitigates truncation artifacts. Compared with a naive domain extension using fixed resolution levels and a fixed sampling rate, the adaptive strategy reduces computational time by over 60% for an image volume of 800x800x600, while preserving the PSNR within the truncated FOV.
Neural Representation-Based Method for Metal-induced Artifact Reduction in Dental CBCT Imaging
Jul 27, 2023Abstract:This study introduces a novel reconstruction method for dental cone-beam computed tomography (CBCT), focusing on effectively reducing metal-induced artifacts commonly encountered in the presence of prevalent metallic implants. Despite significant progress in metal artifact reduction techniques, challenges persist owing to the intricate physical interactions between polychromatic X-ray beams and metal objects, which are further compounded by the additional effects associated with metal-tooth interactions and factors specific to the dental CBCT data environment. To overcome these limitations, we propose an implicit neural network that generates two distinct and informative tomographic images. One image represents the monochromatic attenuation distribution at a specific energy level, whereas the other captures the nonlinear beam-hardening factor resulting from the polychromatic nature of X-ray beams. In contrast to existing CT reconstruction techniques, the proposed method relies exclusively on the Beer--Lambert law, effectively preventing the generation of metal-induced artifacts during the backprojection process commonly implemented in conventional methods. Extensive experimental evaluations demonstrate that the proposed method effectively reduces metal artifacts while providing high-quality image reconstructions, thus emphasizing the significance of the second image in capturing the nonlinear beam-hardening factor.
A robust multi-domain network for short-scanning amyloid PET reconstruction
May 17, 2023



Abstract:This paper presents a robust multi-domain network designed to restore low-quality amyloid PET images acquired in a short period of time. The proposed method is trained on pairs of PET images from short (2 minutes) and standard (20 minutes) scanning times, sourced from multiple domains. Learning relevant image features between these domains with a single network is challenging. Our key contribution is the introduction of a mapping label, which enables effective learning of specific representations between different domains. The network, trained with various mapping labels, can efficiently correct amyloid PET datasets in multiple training domains and unseen domains, such as those obtained with new radiotracers, acquisition protocols, or PET scanners. Internal, temporal, and external validations demonstrate the effectiveness of the proposed method. Notably, for external validation datasets from unseen domains, the proposed method achieved comparable or superior results relative to methods trained with these datasets, in terms of quantitative metrics such as normalized root mean-square error and structure similarity index measure. Two nuclear medicine physicians evaluated the amyloid status as positive or negative for the external validation datasets, with accuracies of 0.970 and 0.930 for readers 1 and 2, respectively.
Automatic 3D Registration of Dental CBCT and Face Scan Data using 2D Projection images
May 17, 2023



Abstract:This paper presents a fully automatic registration method of dental cone-beam computed tomography (CBCT) and face scan data. It can be used for a digital platform of 3D jaw-teeth-face models in a variety of applications, including 3D digital treatment planning and orthognathic surgery. Difficulties in accurately merging facial scans and CBCT images are due to the different image acquisition methods and limited area of correspondence between the two facial surfaces. In addition, it is difficult to use machine learning techniques because they use face-related 3D medical data with radiation exposure, which are difficult to obtain for training. The proposed method addresses these problems by reusing an existing machine-learning-based 2D landmark detection algorithm in an open-source library and developing a novel mathematical algorithm that identifies paired 3D landmarks from knowledge of the corresponding 2D landmarks. A main contribution of this study is that the proposed method does not require annotated training data of facial landmarks because it uses a pre-trained facial landmark detection algorithm that is known to be robust and generalized to various 2D face image models. Note that this reduces a 3D landmark detection problem to a 2D problem of identifying the corresponding landmarks on two 2D projection images generated from two different projection angles. Here, the 3D landmarks for registration were selected from the sub-surfaces with the least geometric change under the CBCT and face scan environments. For the final fine-tuning of the registration, the Iterative Closest Point method was applied, which utilizes geometrical information around the 3D landmarks. The experimental results show that the proposed method achieved an averaged surface distance error of 0.74 mm for three pairs of CBCT and face scan datasets.
Machine Learning-based Signal Quality Assessment for Cardiac Volume Monitoring in Electrical Impedance Tomography
Jan 04, 2023Abstract:Owing to recent advances in thoracic electrical impedance tomography, a patient's hemodynamic function can be noninvasively and continuously estimated in real-time by surveilling a cardiac volume signal associated with stroke volume and cardiac output. In clinical applications, however, a cardiac volume signal is often of low quality, mainly because of the patient's deliberate movements or inevitable motions during clinical interventions. This study aims to develop a signal quality indexing method that assesses the influence of motion artifacts on transient cardiac volume signals. The assessment is performed on each cardiac cycle to take advantage of the periodicity and regularity in cardiac volume changes. Time intervals are identified using the synchronized electrocardiography system. We apply divergent machine-learning methods, which can be sorted into discriminative-model and manifold-learning approaches. The use of machine-learning could be suitable for our real-time monitoring application that requires fast inference and automation as well as high accuracy. In the clinical environment, the proposed method can be utilized to provide immediate warnings so that clinicians can minimize confusion regarding patients' conditions, reduce clinical resource utilization, and improve the confidence level of the monitoring system. Numerous experiments using actual EIT data validate the capability of cardiac volume signals degraded by motion artifacts to be accurately and automatically assessed in real-time by machine learning. The best model achieved an accuracy of 0.95, positive and negative predictive values of 0.96 and 0.86, sensitivity of 0.98, specificity of 0.77, and AUC of 0.96.
Automatic Three-Dimensional Cephalometric Annotation System Using Three-Dimensional Convolutional Neural Networks
Nov 19, 2018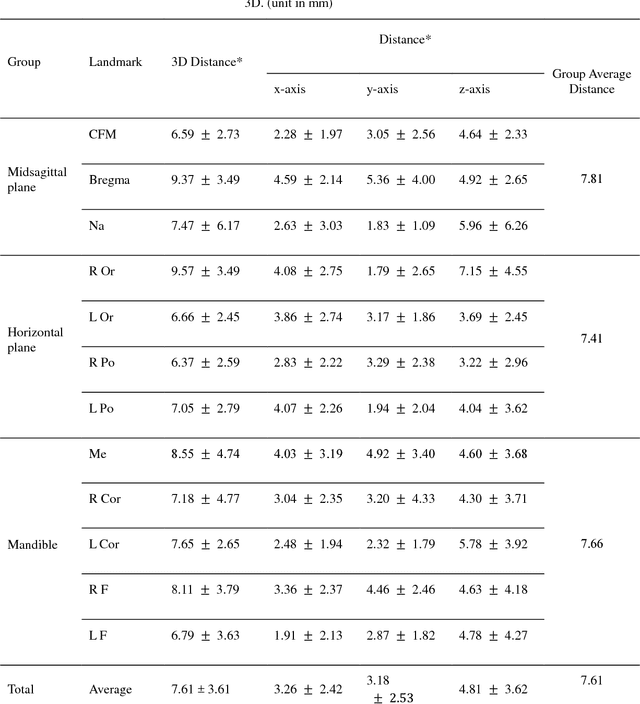



Abstract:Background: Three-dimensional (3D) cephalometric analysis using computerized tomography data has been rapidly adopted for dysmorphosis and anthropometry. Several different approaches to automatic 3D annotation have been proposed to overcome the limitations of traditional cephalometry. The purpose of this study was to evaluate the accuracy of our newly-developed system using a deep learning algorithm for automatic 3D cephalometric annotation. Methods: To overcome current technical limitations, some measures were developed to directly annotate 3D human skull data. Our deep learning-based model system mainly consisted of a 3D convolutional neural network and image data resampling. Results: The discrepancies between the referenced and predicted coordinate values in three axes and in 3D distance were calculated to evaluate system accuracy. Our new model system yielded prediction errors of 3.26, 3.18, and 4.81 mm (for three axes) and 7.61 mm (for 3D). Moreover, there was no difference among the landmarks of the three groups, including the midsagittal plane, horizontal plane, and mandible (p>0.05). Conclusion: A new 3D convolutional neural network-based automatic annotation system for 3D cephalometry was developed. The strategies used to implement the system were detailed and measurement results were evaluated for accuracy. Further development of this system is planned for full clinical application of automatic 3D cephalometric annotation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge