Irene Balelli
EPIONE, UCA
Fed-BioMed: Open, Transparent and Trusted Federated Learning for Real-world Healthcare Applications
Apr 24, 2023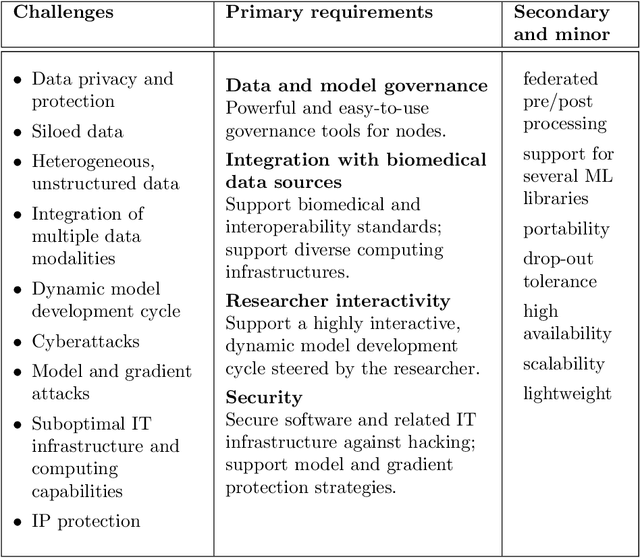
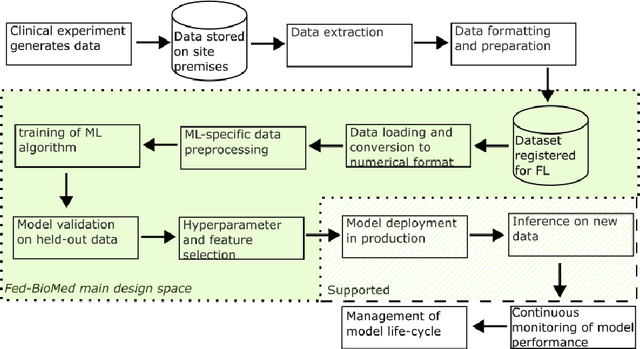
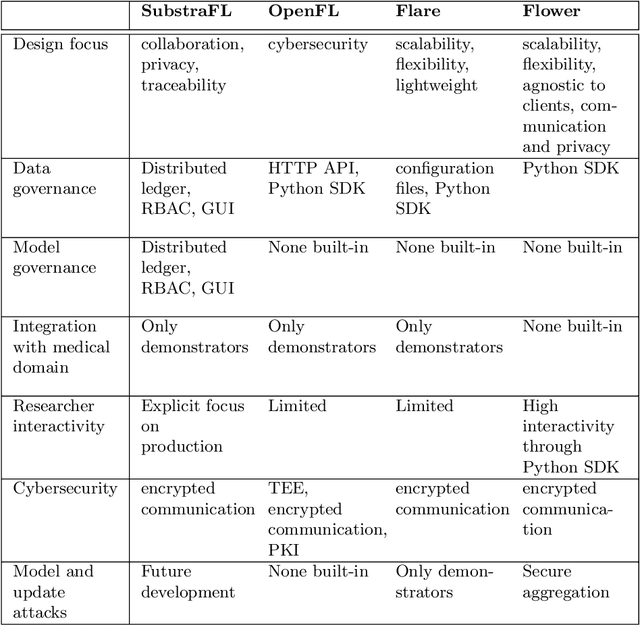
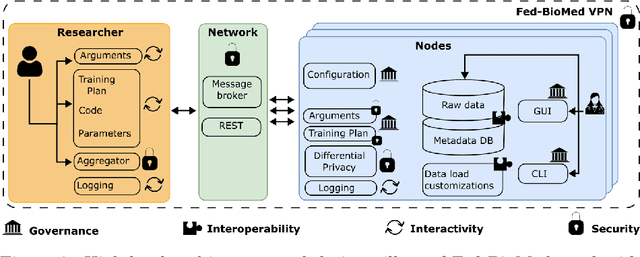
Abstract:The real-world implementation of federated learning is complex and requires research and development actions at the crossroad between different domains ranging from data science, to software programming, networking, and security. While today several FL libraries are proposed to data scientists and users, most of these frameworks are not designed to find seamless application in medical use-cases, due to the specific challenges and requirements of working with medical data and hospital infrastructures. Moreover, governance, design principles, and security assumptions of these frameworks are generally not clearly illustrated, thus preventing the adoption in sensitive applications. Motivated by the current technological landscape of FL in healthcare, in this document we present Fed-BioMed: a research and development initiative aiming at translating federated learning (FL) into real-world medical research applications. We describe our design space, targeted users, domain constraints, and how these factors affect our current and future software architecture.
Fed-MIWAE: Federated Imputation of Incomplete Data via Deep Generative Models
Apr 17, 2023

Abstract:Federated learning allows for the training of machine learning models on multiple decentralized local datasets without requiring explicit data exchange. However, data pre-processing, including strategies for handling missing data, remains a major bottleneck in real-world federated learning deployment, and is typically performed locally. This approach may be biased, since the subpopulations locally observed at each center may not be representative of the overall one. To address this issue, this paper first proposes a more consistent approach to data standardization through a federated model. Additionally, we propose Fed-MIWAE, a federated version of the state-of-the-art imputation method MIWAE, a deep latent variable model for missing data imputation based on variational autoencoders. MIWAE has the great advantage of being easily trainable with classical federated aggregators. Furthermore, it is able to deal with MAR (Missing At Random) data, a more challenging missing-data mechanism than MCAR (Missing Completely At Random), where the missingness of a variable can depend on the observed ones. We evaluate our method on multi-modal medical imaging data and clinical scores from a simulated federated scenario with the ADNI dataset. We compare Fed-MIWAE with respect to classical imputation methods, either performed locally or in a centralized fashion. Fed-MIWAE allows to achieve imputation accuracy comparable with the best centralized method, even when local data distributions are highly heterogeneous. In addition, thanks to the variational nature of Fed-MIWAE, our method is designed to perform multiple imputation, allowing for the quantification of the imputation uncertainty in the federated scenario.
A Differentially Private Probabilistic Framework for Modeling the Variability Across Federated Datasets of Heterogeneous Multi-View Observations
Apr 26, 2022
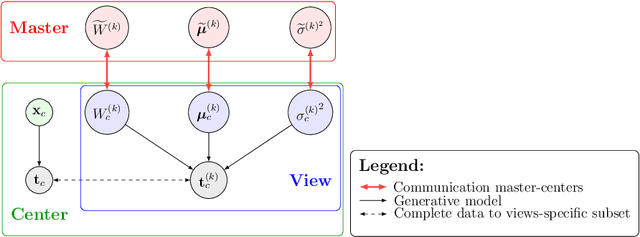
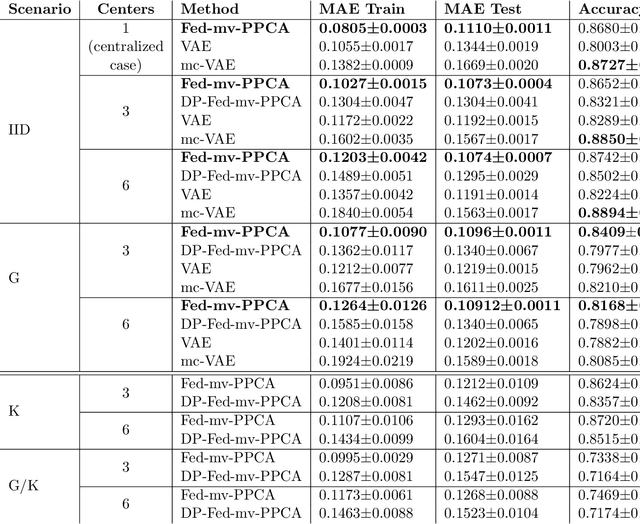

Abstract:We propose a novel federated learning paradigm to model data variability among heterogeneous clients in multi-centric studies. Our method is expressed through a hierarchical Bayesian latent variable model, where client-specific parameters are assumed to be realization from a global distribution at the master level, which is in turn estimated to account for data bias and variability across clients. We show that our framework can be effectively optimized through expectation maximization (EM) over latent master's distribution and clients' parameters. We also introduce formal differential privacy (DP) guarantees compatibly with our EM optimization scheme. We tested our method on the analysis of multi-modal medical imaging data and clinical scores from distributed clinical datasets of patients affected by Alzheimer's disease. We demonstrate that our method is robust when data is distributed either in iid and non-iid manners, even when local parameters perturbation is included to provide DP guarantees. Moreover, the variability of data, views and centers can be quantified in an interpretable manner, while guaranteeing high-quality data reconstruction as compared to state-of-the-art autoencoding models and federated learning schemes. The code is available at https://gitlab.inria.fr/epione/federated-multi-views-ppca.
* Accepted for publication at the Journal of Machine Learning for Biomedical Imaging (MELBA) https://www.melba-journal.org
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge