Haiyan Cen
College of Biosystems Engineering and Food Science, Zhejiang University, Hangzhou, P.R. China, Key Laboratory of Spectroscopy Sensing, Ministry of Agriculture and Rural Affairs, Hangzhou, P.R. China
Real-time Reflectance Generation for UAV Multispectral Imagery using an Onboard Downwelling Spectrometer in Varied Weather Conditions
Dec 27, 2024



Abstract:Advancements in unmanned aerial vehicle (UAV) remote sensing with spectral imaging enable efficient assessment of critical agronomic traits. However, existing reflectance calibration or generation methods suffer from limited prediction accuracy and practical flexibility. This study explores reliable and cost-efficient methods for the accurate conversion of digital number values acquired from a multispectral imager into reflectance, leveraging real-time solar spectra as references. To ensure consistent measurements of incident light, an upward gimbal-mounted downwelling spectrometer was attached to the UAV, and a sinusoidal model was developed to correct for solar position variability. Using principal component analysis on the reference solar spectrum for band selection, a multiple linear regression model with four sensitive bands (4-Band MLR) and a 30 nm bandwidth achieved performance comparable to the direct correction method. The root mean square error (RMSE) for reflectance prediction improved by 86.1% compared to the empirical line method under fluctuating cloudy conditions and by 59.6% compared to the downwelling light sensor method averaged across different weather conditions. The RMSE was calculated as 2.24% in a ground-based diurnal validation, and 2.03% in a UAV campaign conducted at various times throughout a sunny day. Implementing the 4-Band MLR model enhanced the consistency of canopy reflectance within a homogeneous vegetation area by 95.0% during spectral imaging in a large rice field under significant cloud fluctuations. Additionally, improvements of 86.0% and 90.3% were noted for two vegetation indices: the normalized difference vegetation index (NDVI; a ratio index) and the difference vegetation index (DVI; a non-ratio index), respectively.
Biomass phenotyping of oilseed rape through UAV multi-view oblique imaging with 3DGS and SAM model
Nov 13, 2024



Abstract:Biomass estimation of oilseed rape is crucial for optimizing crop productivity and breeding strategies. While UAV-based imaging has advanced high-throughput phenotyping, current methods often rely on orthophoto images, which struggle with overlapping leaves and incomplete structural information in complex field environments. This study integrates 3D Gaussian Splatting (3DGS) with the Segment Anything Model (SAM) for precise 3D reconstruction and biomass estimation of oilseed rape. UAV multi-view oblique images from 36 angles were used to perform 3D reconstruction, with the SAM module enhancing point cloud segmentation. The segmented point clouds were then converted into point cloud volumes, which were fitted to ground-measured biomass using linear regression. The results showed that 3DGS (7k and 30k iterations) provided high accuracy, with peak signal-to-noise ratios (PSNR) of 27.43 and 29.53 and training times of 7 and 49 minutes, respectively. This performance exceeded that of structure from motion (SfM) and mipmap Neural Radiance Fields (Mip-NeRF), demonstrating superior efficiency. The SAM module achieved high segmentation accuracy, with a mean intersection over union (mIoU) of 0.961 and an F1-score of 0.980. Additionally, a comparison of biomass extraction models found the point cloud volume model to be the most accurate, with an determination coefficient (R2) of 0.976, root mean square error (RMSE) of 2.92 g/plant, and mean absolute percentage error (MAPE) of 6.81%, outperforming both the plot crop volume and individual crop volume models. This study highlights the potential of combining 3DGS with multi-view UAV imaging for improved biomass phenotyping.
PanicleNeRF: low-cost, high-precision in-field phenotypingof rice panicles with smartphone
Aug 04, 2024Abstract:The rice panicle traits significantly influence grain yield, making them a primary target for rice phenotyping studies. However, most existing techniques are limited to controlled indoor environments and difficult to capture the rice panicle traits under natural growth conditions. Here, we developed PanicleNeRF, a novel method that enables high-precision and low-cost reconstruction of rice panicle three-dimensional (3D) models in the field using smartphone. The proposed method combined the large model Segment Anything Model (SAM) and the small model You Only Look Once version 8 (YOLOv8) to achieve high-precision segmentation of rice panicle images. The NeRF technique was then employed for 3D reconstruction using the images with 2D segmentation. Finally, the resulting point clouds are processed to successfully extract panicle traits. The results show that PanicleNeRF effectively addressed the 2D image segmentation task, achieving a mean F1 Score of 86.9% and a mean Intersection over Union (IoU) of 79.8%, with nearly double the boundary overlap (BO) performance compared to YOLOv8. As for point cloud quality, PanicleNeRF significantly outperformed traditional SfM-MVS (structure-from-motion and multi-view stereo) methods, such as COLMAP and Metashape. The panicle length was then accurately extracted with the rRMSE of 2.94% for indica and 1.75% for japonica rice. The panicle volume estimated from 3D point clouds strongly correlated with the grain number (R2 = 0.85 for indica and 0.82 for japonica) and grain mass (0.80 for indica and 0.76 for japonica). This method provides a low-cost solution for high-throughput in-field phenotyping of rice panicles, accelerating the efficiency of rice breeding.
Investigation on data fusion of sun-induced chlorophyll fluorescence and reflectance for photosynthetic capacity of rice
Dec 01, 2023Abstract:Studying crop photosynthesis is crucial for improving yield, but current methods are labor-intensive. This research aims to enhance accuracy by combining leaf reflectance and sun-induced chlorophyll fluorescence (SIF) signals to estimate key photosynthetic traits in rice. The study analyzes 149 leaf samples from two rice cultivars, considering reflectance, SIF, chlorophyll, carotenoids, and CO2 response curves. After noise removal, SIF and reflectance spectra are used for data fusion at different levels (raw, feature, and decision). Competitive adaptive reweighted sampling (CARS) extracts features, and partial least squares regression (PLSR) builds regression models. Results indicate that using either reflectance or SIF alone provides modest estimations for photosynthetic traits. However, combining these data sources through measurement-level data fusion significantly improves accuracy, with mid-level and decision-level fusion also showing positive outcomes. In particular, decision-level fusion enhances predictive capabilities, suggesting the potential for efficient crop phenotyping. Overall, sun-induced chlorophyll fluorescence spectra effectively predict rice's photosynthetic capacity, and data fusion methods contribute to increased accuracy, paving the way for high-throughput crop phenotyping.
Generating high-quality 3DMPCs by adaptive data acquisition and NeREF-based reflectance correction to facilitate efficient plant phenotyping
May 11, 2023



Abstract:Non-destructive assessments of plant phenotypic traits using high-quality three-dimensional (3D) and multispectral data can deepen breeders' understanding of plant growth and allow them to make informed managerial decisions. However, subjective viewpoint selection and complex illumination effects under natural light conditions decrease the data quality and increase the difficulty of resolving phenotypic parameters. We proposed methods for adaptive data acquisition and reflectance correction respectively, to generate high-quality 3D multispectral point clouds (3DMPCs) of plants. In the first stage, we proposed an efficient next-best-view (NBV) planning method based on a novel UGV platform with a multi-sensor-equipped robotic arm. In the second stage, we eliminated the illumination effects by using the neural reference field (NeREF) to predict the digital number (DN) of the reference. We tested them on 6 perilla and 6 tomato plants, and selected 2 visible leaves and 4 regions of interest (ROIs) for each plant to assess the biomass and the chlorophyll content. For NBV planning, the average execution time for single perilla and tomato plant at a joint speed of 1.55 rad/s was 58.70 s and 53.60 s respectively. The whole-plant data integrity was improved by an average of 27% compared to using fixed viewpoints alone, and the coefficients of determination (R2) for leaf biomass estimation reached 0.99 and 0.92. For reflectance correction, the average root mean squared error of the reflectance spectra with hemisphere reference-based correction at different ROIs was 0.08 and 0.07 for perilla and tomato. The R2 of chlorophyll content estimation was 0.91 and 0.93 respectively when principal component analysis and Gaussian process regression were applied. Our approach is promising for generating high-quality 3DMPCs of plants under natural light conditions and facilitates accurate plant phenotyping.
PST: Plant Segmentation Transformer Enhanced Phenotyping of MLS Oilseed Rape Point Cloud
Jun 27, 2022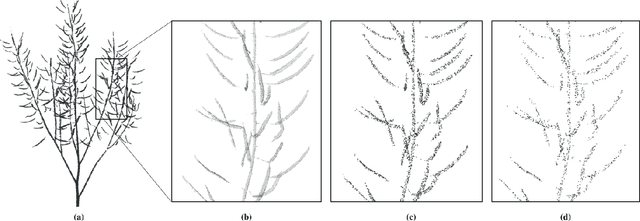
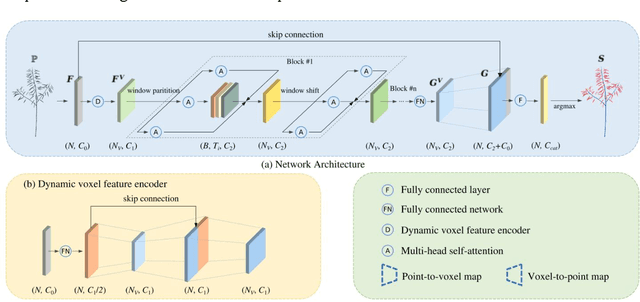
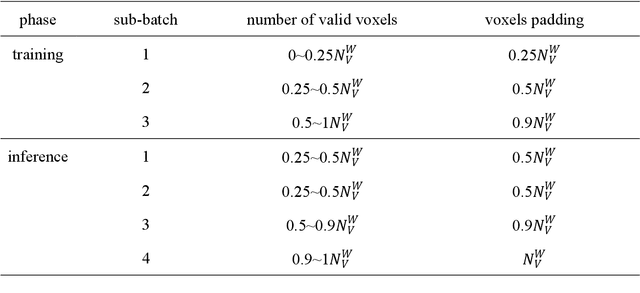
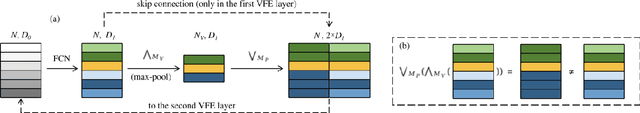
Abstract:Segmentation of plant point clouds to obtain high-precise morphological traits is essential for plant phenotyping and crop breeding. Although the bloom of deep learning methods has boosted much research on the segmentation of plant point cloud, most works follow the common practice of hard voxelization-based or down-sampling-based methods. They are limited to segmenting simple plant organs, overlooking the difficulties of resolving complex plant point clouds with high spatial resolution. In this study, we propose a deep learning network plant segmentation transformer (PST) to realize the semantic and instance segmentation of MLS (Mobile Laser Scanning) oilseed rape point cloud, which characterizes tiny siliques and dense points as the main traits targeted. PST is composed of: (i) a dynamic voxel feature encoder (DVFE) to aggregate per point features with raw spatial resolution; (ii) dual window sets attention block to capture the contextual information; (iii) a dense feature propagation module to obtain the final dense point feature map. The results proved that PST and PST-PointGroup (PG) achieved state-of-the-art performance in semantic and instance segmentation tasks. For semantic segmentation, PST reached 93.96%, 97.29%, 96.52%, 96.88%, and 97.07% in mean IoU, mean Precision, mean Recall, mean F1-score, and overall accuracy, respectively. For instance segmentation, PST-PG reached 89.51%, 89.85%, 88.83% and 82.53% in mCov, mWCov, mPerc90, and mRec90, respectively. This study extends the phenotyping of oilseed rape in an end-to-end way and proves that the deep learning method has a great potential for understanding dense plant point clouds with complex morphological traits.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge