Fiona Inglis
InstanSeg: an embedding-based instance segmentation algorithm optimized for accurate, efficient and portable cell segmentation
Aug 28, 2024
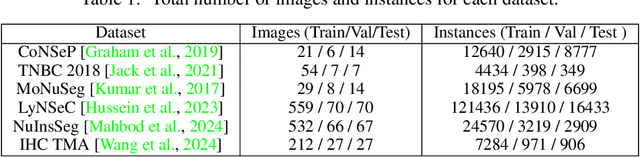

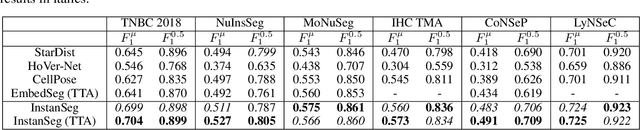
Abstract:Cell and nucleus segmentation are fundamental tasks for quantitative bioimage analysis. Despite progress in recent years, biologists and other domain experts still require novel algorithms to handle increasingly large and complex real-world datasets. These algorithms must not only achieve state-of-the-art accuracy, but also be optimized for efficiency, portability and user-friendliness. Here, we introduce InstanSeg: a novel embedding-based instance segmentation pipeline designed to identify cells and nuclei in microscopy images. Using six public cell segmentation datasets, we demonstrate that InstanSeg can significantly improve accuracy when compared to the most widely used alternative methods, while reducing the processing time by at least 60%. Furthermore, InstanSeg is designed to be fully serializable as TorchScript and supports GPU acceleration on a range of hardware. We provide an open-source implementation of InstanSeg in Python, in addition to a user-friendly, interactive QuPath extension for inference written in Java. Our code and pre-trained models are available at https://github.com/instanseg/instanseg .
Open and reusable deep learning for pathology with WSInfer and QuPath
Sep 08, 2023Abstract:The field of digital pathology has seen a proliferation of deep learning models in recent years. Despite substantial progress, it remains rare for other researchers and pathologists to be able to access models published in the literature and apply them to their own images. This is due to difficulties in both sharing and running models. To address these concerns, we introduce WSInfer: a new, open-source software ecosystem designed to make deep learning for pathology more streamlined and accessible. WSInfer comprises three main elements: 1) a Python package and command line tool to efficiently apply patch-based deep learning inference to whole slide images; 2) a QuPath extension that provides an alternative inference engine through user-friendly and interactive software, and 3) a model zoo, which enables pathology models and metadata to be easily shared in a standardized form. Together, these contributions aim to encourage wider reuse, exploration, and interrogation of deep learning models for research purposes, by putting them into the hands of pathologists and eliminating a need for coding experience when accessed through QuPath. The WSInfer source code is hosted on GitHub and documentation is available at https://wsinfer.readthedocs.io.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge