Douglas E. V. Pires
The University of Melbourne
Explainable Machine Learning for ICU Readmission Prediction
Sep 27, 2023



Abstract:The intensive care unit (ICU) comprises a complex hospital environment, where decisions made by clinicians have a high level of risk for the patients' lives. A comprehensive care pathway must then be followed to reduce p complications. Uncertain, competing and unplanned aspects within this environment increase the difficulty in uniformly implementing the care pathway. Readmission contributes to this pathway's difficulty, occurring when patients are admitted again to the ICU in a short timeframe, resulting in high mortality rates and high resource utilisation. Several works have tried to predict readmission through patients' medical information. Although they have some level of success while predicting readmission, those works do not properly assess, characterise and understand readmission prediction. This work proposes a standardised and explainable machine learning pipeline to model patient readmission on a multicentric database (i.e., the eICU cohort with 166,355 patients, 200,859 admissions and 6,021 readmissions) while validating it on monocentric (i.e., the MIMIC IV cohort with 382,278 patients, 523,740 admissions and 5,984 readmissions) and multicentric settings. Our machine learning pipeline achieved predictive performance in terms of the area of the receiver operating characteristic curve (AUC) up to 0.7 with a Random Forest classification model, yielding an overall good calibration and consistency on validation sets. From explanations provided by the constructed models, we could also derive a set of insightful conclusions, primarily on variables related to vital signs and blood tests (e.g., albumin, blood urea nitrogen and hemoglobin levels), demographics (e.g., age, and admission height and weight), and ICU-associated variables (e.g., unit type). These insights provide an invaluable source of information during clinicians' decision-making while discharging ICU patients.
AI driven B-cell Immunotherapy Design
Sep 03, 2023Abstract:Antibodies, a prominent class of approved biologics, play a crucial role in detecting foreign antigens. The effectiveness of antigen neutralisation and elimination hinges upon the strength, sensitivity, and specificity of the paratope-epitope interaction, which demands resource-intensive experimental techniques for characterisation. In recent years, artificial intelligence and machine learning methods have made significant strides, revolutionising the prediction of protein structures and their complexes. The past decade has also witnessed the evolution of computational approaches aiming to support immunotherapy design. This review focuses on the progress of machine learning-based tools and their frameworks in the domain of B-cell immunotherapy design, encompassing linear and conformational epitope prediction, paratope prediction, and antibody design. We mapped the most commonly used data sources, evaluation metrics, and method availability and thoroughly assessed their significance and limitations, discussing the main challenges ahead.
Large-scale protein-protein post-translational modification extraction with distant supervision and confidence calibrated BioBERT
Jan 06, 2022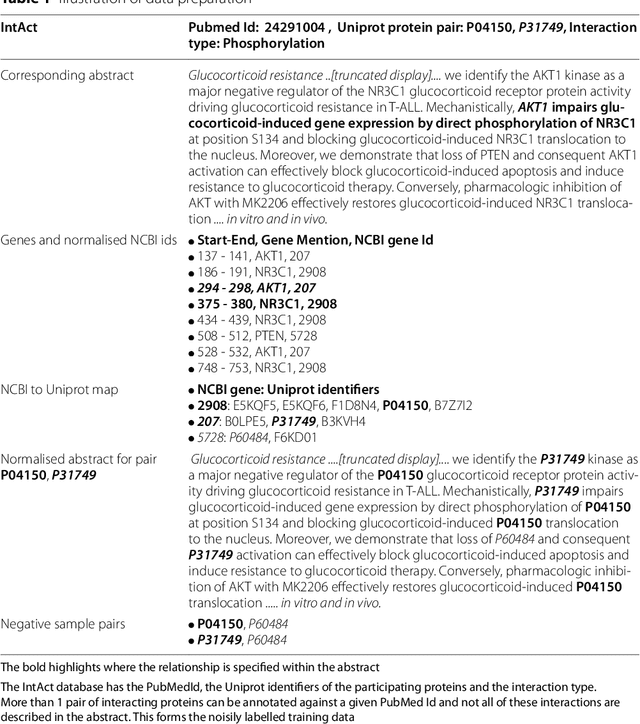


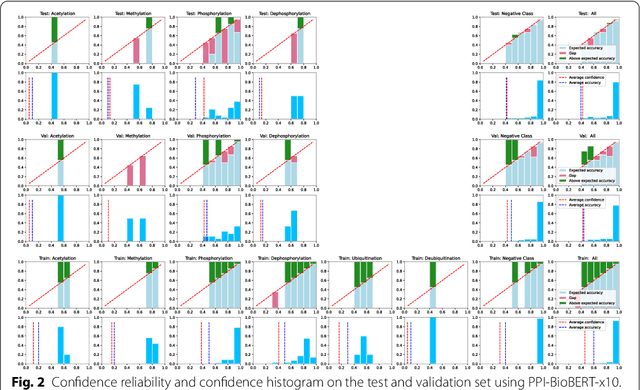
Abstract:Protein-protein interactions (PPIs) are critical to normal cellular function and are related to many disease pathways. However, only 4% of PPIs are annotated with PTMs in biological knowledge databases such as IntAct, mainly performed through manual curation, which is neither time nor cost-effective. We use the IntAct PPI database to create a distant supervised dataset annotated with interacting protein pairs, their corresponding PTM type, and associated abstracts from the PubMed database. We train an ensemble of BioBERT models - dubbed PPI-BioBERT-x10 to improve confidence calibration. We extend the use of ensemble average confidence approach with confidence variation to counteract the effects of class imbalance to extract high confidence predictions. The PPI-BioBERT-x10 model evaluated on the test set resulted in a modest F1-micro 41.3 (P =5 8.1, R = 32.1). However, by combining high confidence and low variation to identify high quality predictions, tuning the predictions for precision, we retained 19% of the test predictions with 100% precision. We evaluated PPI-BioBERT-x10 on 18 million PubMed abstracts and extracted 1.6 million (546507 unique PTM-PPI triplets) PTM-PPI predictions, and filter ~ 5700 (4584 unique) high confidence predictions. Of the 5700, human evaluation on a small randomly sampled subset shows that the precision drops to 33.7% despite confidence calibration and highlights the challenges of generalisability beyond the test set even with confidence calibration. We circumvent the problem by only including predictions associated with multiple papers, improving the precision to 58.8%. In this work, we highlight the benefits and challenges of deep learning-based text mining in practice, and the need for increased emphasis on confidence calibration to facilitate human curation efforts.
* BMC BioInformatics
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge