David B. Ascher
The University of Queensland
Auto-ADMET: An Effective and Interpretable AutoML Method for Chemical ADMET Property Prediction
Feb 22, 2025



Abstract:Machine learning (ML) has been playing important roles in drug discovery in the past years by providing (pre-)screening tools for prioritising chemical compounds to pass through wet lab experiments. One of the main ML tasks in drug discovery is to build quantitative structure-activity relationship (QSAR) models, associating the molecular structure of chemical compounds with an activity or property. These properties -- including absorption, distribution, metabolism, excretion and toxicity (ADMET) -- are essential to model compound behaviour, activity and interactions in the organism. Although several methods exist, the majority of them do not provide an appropriate model's personalisation, yielding to bias and lack of generalisation to new data since the chemical space usually shifts from application to application. This fact leads to low predictive performance when completely new data is being tested by the model. The area of Automated Machine Learning (AutoML) emerged aiming to solve this issue, outputting tailored ML algorithms to the data at hand. Although an important task, AutoML has not been practically used to assist cheminformatics and computational chemistry researchers often, with just a few works related to the field. To address these challenges, this work introduces Auto-ADMET, an interpretable evolutionary-based AutoML method for chemical ADMET property prediction. Auto-ADMET employs a Grammar-based Genetic Programming (GGP) method with a Bayesian Network Model to achieve comparable or better predictive performance against three alternative methods -- standard GGP method, pkCSM and XGBOOST model -- on 12 benchmark chemical ADMET property prediction datasets. The use of a Bayesian Network model on Auto-ADMET's evolutionary process assisted in both shaping the search procedure and interpreting the causes of its AutoML performance.
Towards Evolutionary-based Automated Machine Learning for Small Molecule Pharmacokinetic Prediction
Aug 01, 2024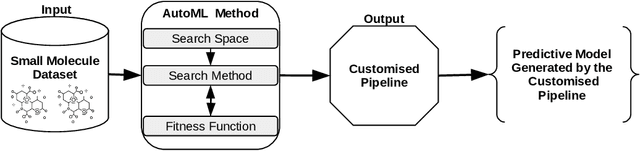
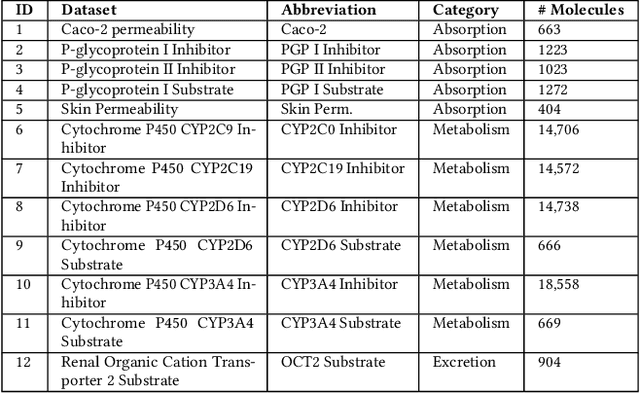
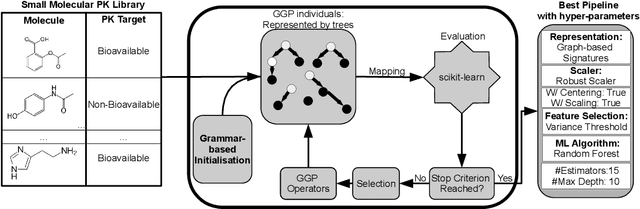
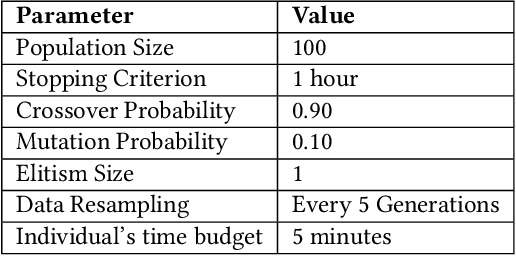
Abstract:Machine learning (ML) is revolutionising drug discovery by expediting the prediction of small molecule properties essential for developing new drugs. These properties -- including absorption, distribution, metabolism and excretion (ADME)-- are crucial in the early stages of drug development since they provide an understanding of the course of the drug in the organism, i.e., the drug's pharmacokinetics. However, existing methods lack personalisation and rely on manually crafted ML algorithms or pipelines, which can introduce inefficiencies and biases into the process. To address these challenges, we propose a novel evolutionary-based automated ML method (AutoML) specifically designed for predicting small molecule properties, with a particular focus on pharmacokinetics. Leveraging the advantages of grammar-based genetic programming, our AutoML method streamlines the process by automatically selecting algorithms and designing predictive pipelines tailored to the particular characteristics of input molecular data. Results demonstrate AutoML's effectiveness in selecting diverse ML algorithms, resulting in comparable or even improved predictive performances compared to conventional approaches. By offering personalised ML-driven pipelines, our method promises to enhance small molecule research in drug discovery, providing researchers with a valuable tool for accelerating the development of novel therapeutic drugs.
Explainable Machine Learning for ICU Readmission Prediction
Sep 27, 2023



Abstract:The intensive care unit (ICU) comprises a complex hospital environment, where decisions made by clinicians have a high level of risk for the patients' lives. A comprehensive care pathway must then be followed to reduce p complications. Uncertain, competing and unplanned aspects within this environment increase the difficulty in uniformly implementing the care pathway. Readmission contributes to this pathway's difficulty, occurring when patients are admitted again to the ICU in a short timeframe, resulting in high mortality rates and high resource utilisation. Several works have tried to predict readmission through patients' medical information. Although they have some level of success while predicting readmission, those works do not properly assess, characterise and understand readmission prediction. This work proposes a standardised and explainable machine learning pipeline to model patient readmission on a multicentric database (i.e., the eICU cohort with 166,355 patients, 200,859 admissions and 6,021 readmissions) while validating it on monocentric (i.e., the MIMIC IV cohort with 382,278 patients, 523,740 admissions and 5,984 readmissions) and multicentric settings. Our machine learning pipeline achieved predictive performance in terms of the area of the receiver operating characteristic curve (AUC) up to 0.7 with a Random Forest classification model, yielding an overall good calibration and consistency on validation sets. From explanations provided by the constructed models, we could also derive a set of insightful conclusions, primarily on variables related to vital signs and blood tests (e.g., albumin, blood urea nitrogen and hemoglobin levels), demographics (e.g., age, and admission height and weight), and ICU-associated variables (e.g., unit type). These insights provide an invaluable source of information during clinicians' decision-making while discharging ICU patients.
AI driven B-cell Immunotherapy Design
Sep 03, 2023Abstract:Antibodies, a prominent class of approved biologics, play a crucial role in detecting foreign antigens. The effectiveness of antigen neutralisation and elimination hinges upon the strength, sensitivity, and specificity of the paratope-epitope interaction, which demands resource-intensive experimental techniques for characterisation. In recent years, artificial intelligence and machine learning methods have made significant strides, revolutionising the prediction of protein structures and their complexes. The past decade has also witnessed the evolution of computational approaches aiming to support immunotherapy design. This review focuses on the progress of machine learning-based tools and their frameworks in the domain of B-cell immunotherapy design, encompassing linear and conformational epitope prediction, paratope prediction, and antibody design. We mapped the most commonly used data sources, evaluation metrics, and method availability and thoroughly assessed their significance and limitations, discussing the main challenges ahead.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge