Cian M. Scannell
Deep learning motion correction of quantitative stress perfusion cardiovascular magnetic resonance
Oct 01, 2025Abstract:Background: Quantitative stress perfusion cardiovascular magnetic resonance (CMR) is a powerful tool for assessing myocardial ischemia. Motion correction is essential for accurate pixel-wise mapping but traditional registration-based methods are slow and sensitive to acquisition variability, limiting robustness and scalability. Methods: We developed an unsupervised deep learning-based motion correction pipeline that replaces iterative registration with efficient one-shot estimation. The method corrects motion in three steps and uses robust principal component analysis to reduce contrast-related effects. It aligns the perfusion series and auxiliary images (arterial input function and proton density-weighted series). Models were trained and validated on multivendor data from 201 patients, with 38 held out for testing. Performance was assessed via temporal alignment and quantitative perfusion values, compared to a previously published registration-based method. Results: The deep learning approach significantly improved temporal smoothness of time-intensity curves (p<0.001). Myocardial alignment (Dice = 0.92 (0.04) and 0.91 (0.05)) was comparable to the baseline and superior to before registration (Dice = 0.80 (0.09), p<0.001). Perfusion maps showed reduced motion, with lower standard deviation in the myocardium (0.52 (0.39) ml/min/g) compared to baseline (0.55 (0.44) ml/min/g). Processing time was reduced 15-fold. Conclusion: This deep learning pipeline enables fast, robust motion correction for stress perfusion CMR, improving accuracy across dynamic and auxiliary images. Trained on multivendor data, it generalizes across sequences and may facilitate broader clinical adoption of quantitative perfusion imaging.
Optimized Automated Cardiac MR Scar Quantification with GAN-Based Data Augmentation
Sep 27, 2021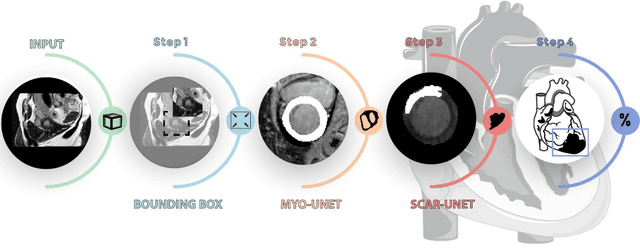
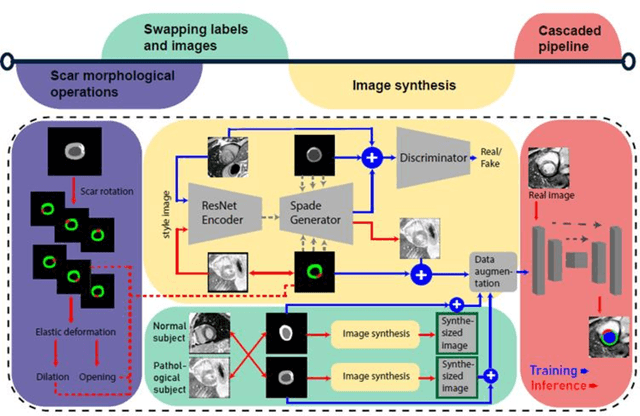
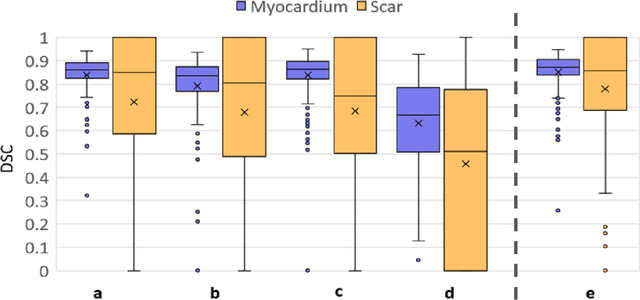
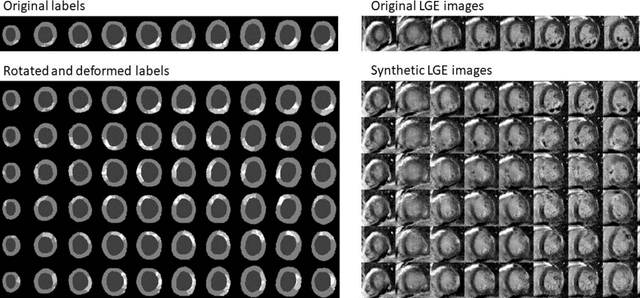
Abstract:Background: The clinical utility of late gadolinium enhancement (LGE) cardiac MRI is limited by the lack of standardization, and time-consuming postprocessing. In this work, we tested the hypothesis that a cascaded deep learning pipeline trained with augmentation by synthetically generated data would improve model accuracy and robustness for automated scar quantification. Methods: A cascaded pipeline consisting of three consecutive neural networks is proposed, starting with a bounding box regression network to identify a region of interest around the left ventricular (LV) myocardium. Two further nnU-Net models are then used to segment the myocardium and, if present, scar. The models were trained on the data from the EMIDEC challenge, supplemented with an extensive synthetic dataset generated with a conditional GAN. Results: The cascaded pipeline significantly outperformed a single nnU-Net directly segmenting both the myocardium (mean Dice similarity coefficient (DSC) (standard deviation (SD)): 0.84 (0.09) vs 0.63 (0.20), p < 0.01) and scar (DSC: 0.72 (0.34) vs 0.46 (0.39), p < 0.01) on a per-slice level. The inclusion of the synthetic data as data augmentation during training improved the scar segmentation DSC by 0.06 (p < 0.01). The mean DSC per-subject on the challenge test set, for the cascaded pipeline augmented by synthetic generated data, was 0.86 (0.03) and 0.67 (0.29) for myocardium and scar, respectively. Conclusion: A cascaded deep learning-based pipeline trained with augmentation by synthetically generated data leads to myocardium and scar segmentations that are similar to the manual operator, and outperforms direct segmentation without the synthetic images.
Physics-informed neural networks for myocardial perfusion MRI quantification
Dec 07, 2020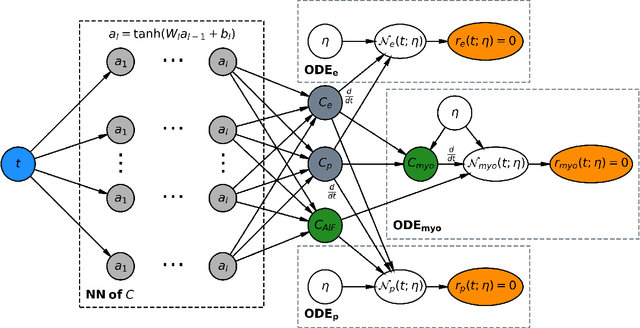
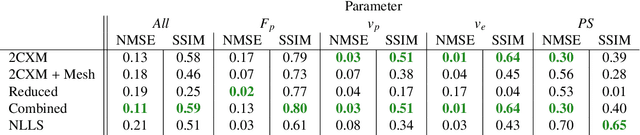
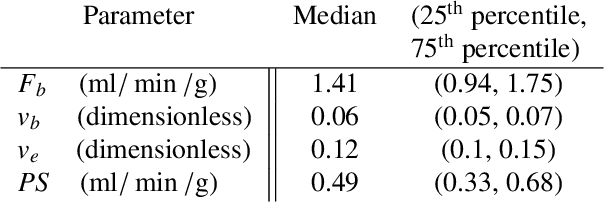
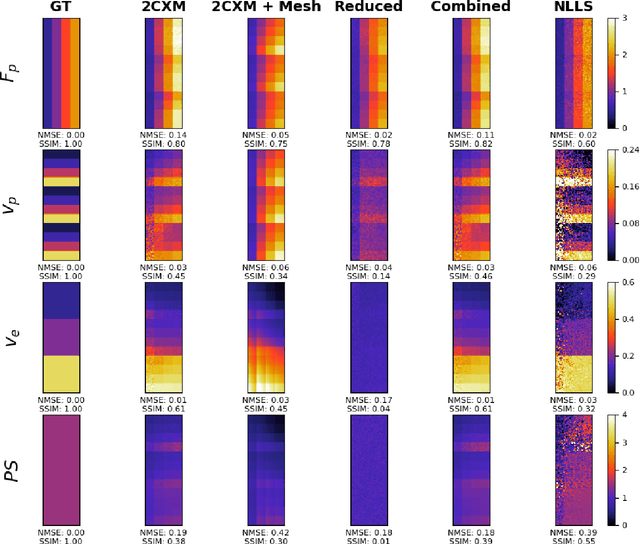
Abstract:Tracer-kinetic models allow for the quantification of kinetic parameters such as blood flow from dynamic contrast-enhanced magnetic resonance (MR) images. Fitting the observed data with multi-compartment exchange models is desirable, as they are physiologically plausible and resolve directly for blood flow and microvascular function. However, the reliability of model fitting is limited by the low signal-to-noise ratio, temporal resolution, and acquisition length. This may result in inaccurate parameter estimates. This study introduces physics-informed neural networks (PINNs) as a means to perform myocardial perfusion MR quantification, which provides a versatile scheme for the inference of kinetic parameters. These neural networks can be trained to fit the observed perfusion MR data while respecting the underlying physical conservation laws described by a multi-compartment exchange model. Here, we provide a framework for the implementation of PINNs in myocardial perfusion MR. The approach is validated both in silico and in vivo. In the in silico study, an overall reduction in mean-squared error with the ground-truth parameters was observed compared to a standard non-linear least squares fitting approach. The in vivo study demonstrates that the method produces parameter values comparable to those previously found in literature, as well as providing parameter maps which match the clinical diagnosis of patients.
Domain-Adversarial Learning for Multi-Centre, Multi-Vendor, and Multi-Disease Cardiac MR Image Segmentation
Aug 26, 2020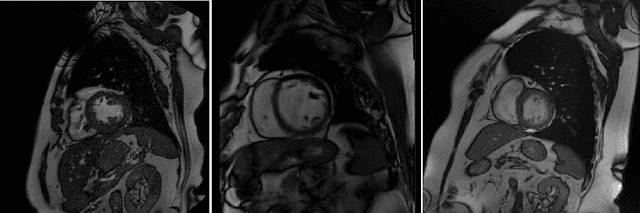
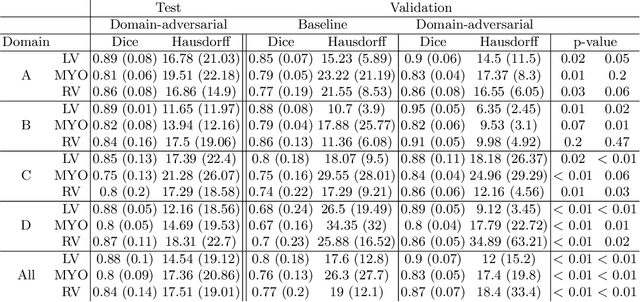
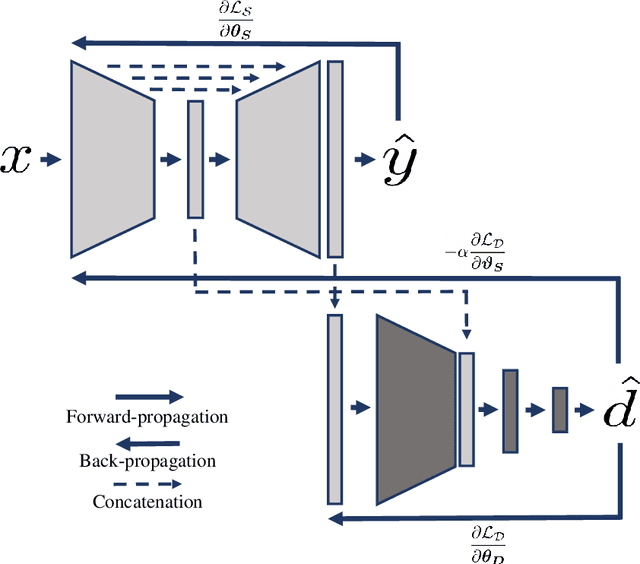
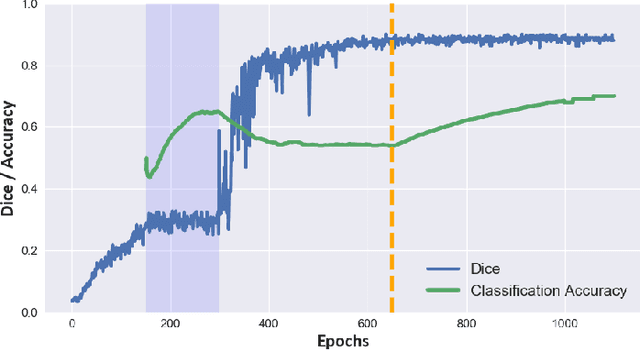
Abstract:Cine cardiac magnetic resonance (CMR) has become the gold standard for the non-invasive evaluation of cardiac function. In particular, it allows the accurate quantification of functional parameters including the chamber volumes and ejection fraction. Deep learning has shown the potential to automate the requisite cardiac structure segmentation. However, the lack of robustness of deep learning models has hindered their widespread clinical adoption. Due to differences in the data characteristics, neural networks trained on data from a specific scanner are not guaranteed to generalise well to data acquired at a different centre or with a different scanner. In this work, we propose a principled solution to the problem of this domain shift. Domain-adversarial learning is used to train a domain-invariant 2D U-Net using labelled and unlabelled data. This approach is evaluated on both seen and unseen domains from the M\&Ms challenge dataset and the domain-adversarial approach shows improved performance as compared to standard training. Additionally, we show that the domain information cannot be recovered from the learned features.
Deep learning-based prediction of kinetic parameters from myocardial perfusion MRI
Jul 27, 2019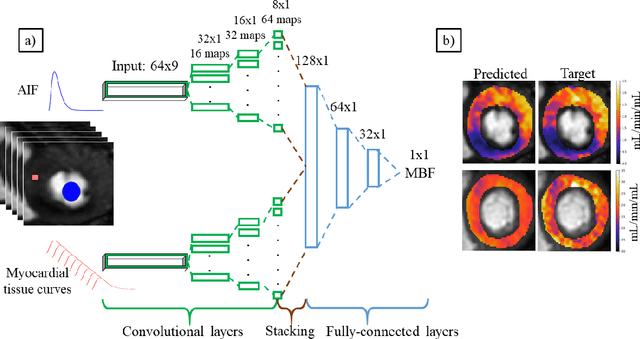
Abstract:The quantification of myocardial perfusion MRI has the potential to provide a fast, automated and user-independent assessment of myocardial ischaemia. However, due to the relatively high noise level and low temporal resolution of the acquired data and the complexity of the tracer-kinetic models, the model fitting can yield unreliable parameter estimates. A solution to this problem is the use of Bayesian inference which can incorporate prior knowledge and improve the reliability of the parameter estimation. This, however, uses Markov chain Monte Carlo sampling to approximate the posterior distribution of the kinetic parameters which is extremely time intensive. This work proposes training convolutional networks to directly predict the kinetic parameters from the signal-intensity curves that are trained using estimates obtained from the Bayesian inference. This allows fast estimation of the kinetic parameters with a similar performance to the Bayesian inference.
Hierarchical Bayesian myocardial perfusion quantification
Jun 06, 2019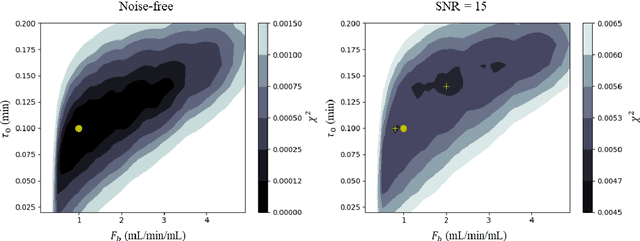
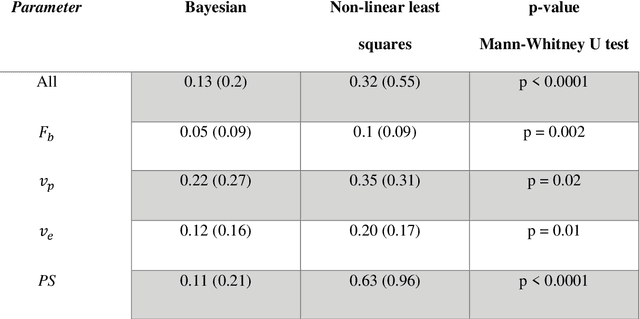
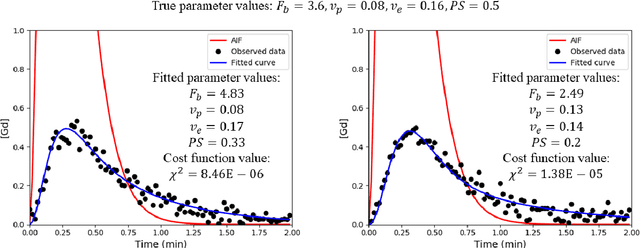
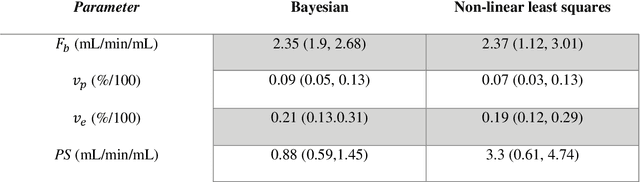
Abstract:Purpose: Tracer-kinetic models can be used for the quantitative assessment of contrast-enhanced MRI data. However, the model-fitting can produce unreliable results due to the limited data acquired and the high noise levels. Such problems are especially prevalent in myocardial perfusion MRI leading to the compromise of constrained numerical deconvolutions and segmental signal averaging being commonly used as alternatives to the more complex tracer-kinetic models. Methods: In this work, the use of hierarchical Bayesian inference for the parameter estimation is explored. It is shown that with Bayesian inference it is possible to reliably fit the two-compartment exchange model to perfusion data. The use of prior knowledge on the ranges of kinetic parameters and the fact that neighbouring voxels are likely to have similar kinetic properties combined with a Markov chain Monte Carlo based fitting procedure significantly improves the reliability of the perfusion estimates with compared to the traditional least-squares approach. The method is assessed using both simulated and patient data. Results: The average (standard deviation) normalised mean square error for the distinct noise realisations of a simulation phantom falls from 0.32 (0.55) with the least-squares fitting to 0.13 (0.2) using Bayesian inference. The assessment of the presence of coronary artery disease based purely on the quantitative MBF maps obtained using Bayesian inference matches the visual assessment in all 24 slices. When using the maps obtained by the least-squares fitting, a corresponding assessment is only achieved in 16/24 slices. Conclusion: Bayesian inference allows a reliable, fully automated and user-independent assessment of myocardial perfusion on a voxel-wise level using the two-compartment exchange model.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge