Chong Yin
MERA: Multimodal and Multiscale Self-Explanatory Model with Considerably Reduced Annotation for Lung Nodule Diagnosis
Apr 27, 2025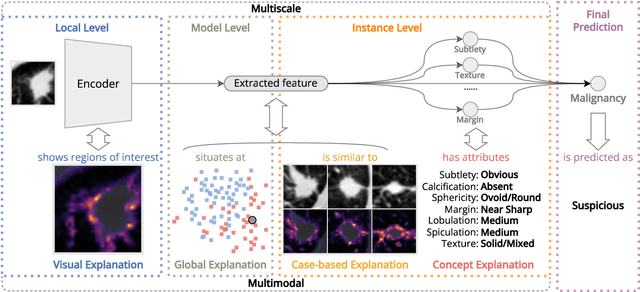
Abstract:Lung cancer, a leading cause of cancer-related deaths globally, emphasises the importance of early detection for better patient outcomes. Pulmonary nodules, often early indicators of lung cancer, necessitate accurate, timely diagnosis. Despite Explainable Artificial Intelligence (XAI) advances, many existing systems struggle providing clear, comprehensive explanations, especially with limited labelled data. This study introduces MERA, a Multimodal and Multiscale self-Explanatory model designed for lung nodule diagnosis with considerably Reduced Annotation requirements. MERA integrates unsupervised and weakly supervised learning strategies (self-supervised learning techniques and Vision Transformer architecture for unsupervised feature extraction) and a hierarchical prediction mechanism leveraging sparse annotations via semi-supervised active learning in the learned latent space. MERA explains its decisions on multiple levels: model-level global explanations via semantic latent space clustering, instance-level case-based explanations showing similar instances, local visual explanations via attention maps, and concept explanations using critical nodule attributes. Evaluations on the public LIDC dataset show MERA's superior diagnostic accuracy and self-explainability. With only 1% annotated samples, MERA achieves diagnostic accuracy comparable to or exceeding state-of-the-art methods requiring full annotation. The model's inherent design delivers comprehensive, robust, multilevel explanations aligned closely with clinical practice, enhancing trustworthiness and transparency. Demonstrated viability of unsupervised and weakly supervised learning lowers the barrier to deploying diagnostic AI in broader medical domains. Our complete code is open-source available: https://github.com/diku-dk/credanno.
cRedAnno+: Annotation Exploitation in Self-Explanatory Lung Nodule Diagnosis
Nov 07, 2022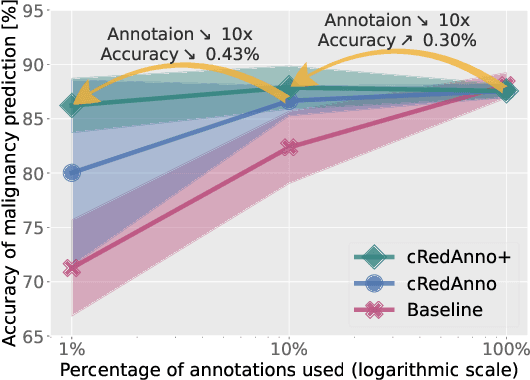
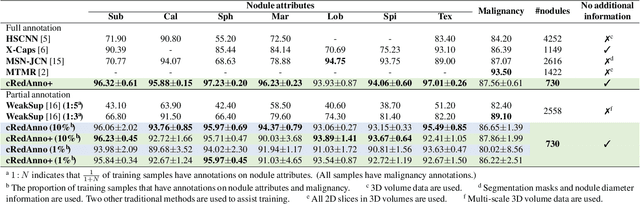
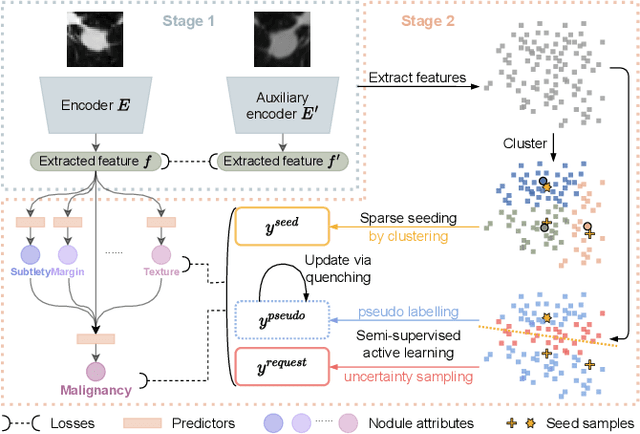
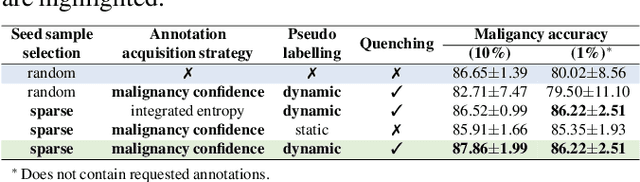
Abstract:Recently, attempts have been made to reduce annotation requirements in feature-based self-explanatory models for lung nodule diagnosis. As a representative, cRedAnno achieves competitive performance with considerably reduced annotation needs by introducing self-supervised contrastive learning to do unsupervised feature extraction. However, it exhibits unstable performance under scarce annotation conditions. To improve the accuracy and robustness of cRedAnno, we propose an annotation exploitation mechanism by conducting semi-supervised active learning with sparse seeding and training quenching in the learned semantically meaningful reasoning space to jointly utilise the extracted features, annotations, and unlabelled data. The proposed approach achieves comparable or even higher malignancy prediction accuracy with 10x fewer annotations, meanwhile showing better robustness and nodule attribute prediction accuracy under the condition of 1% annotations. Our complete code is open-source available: https://github.com/diku-dk/credanno.
Reducing Annotation Need in Self-Explanatory Models for Lung Nodule Diagnosis
Jun 27, 2022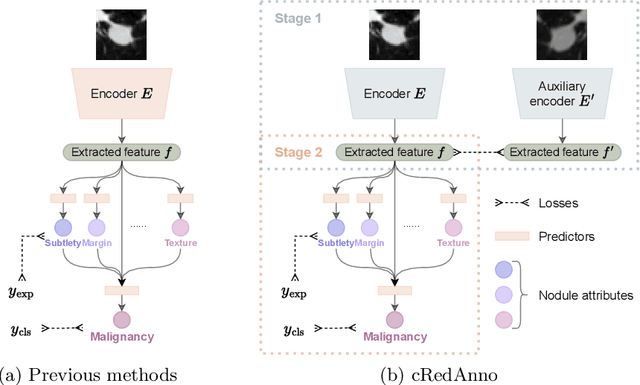
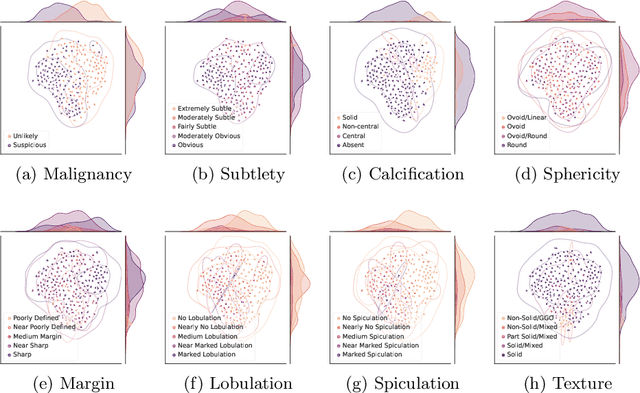
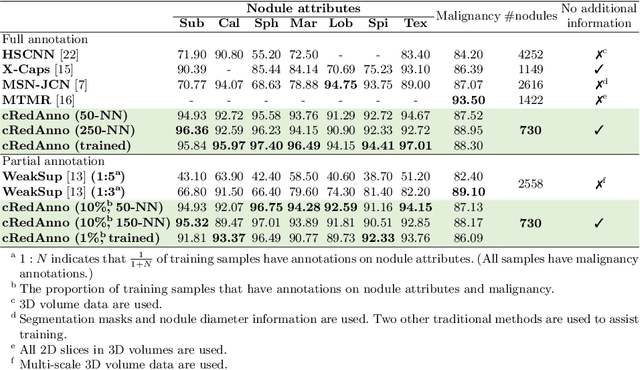
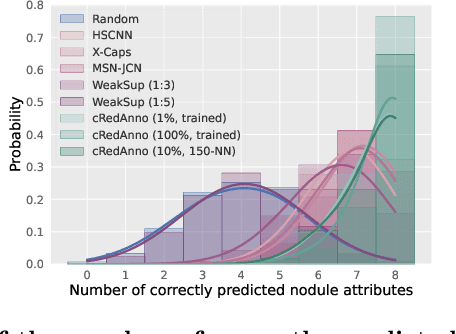

Abstract:Feature-based self-explanatory methods explain their classification in terms of human-understandable features. In the medical imaging community, this semantic matching of clinical knowledge adds significantly to the trustworthiness of the AI. However, the cost of additional annotation of features remains a pressing issue. We address this problem by proposing cRedAnno, a data-/annotation-efficient self-explanatory approach for lung nodule diagnosis. cRedAnno considerably reduces the annotation need by introducing self-supervised contrastive learning to alleviate the burden of learning most parameters from annotation, replacing end-to-end training with two-stage training. When training with hundreds of nodule samples and only 1% of their annotations, cRedAnno achieves competitive accuracy in predicting malignancy, meanwhile significantly surpassing most previous works in predicting nodule attributes. Visualisation of the learned space further indicates that the correlation between the clustering of malignancy and nodule attributes coincides with clinical knowledge. Our complete code is open-source available: https://github.com/ludles/credanno.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge