cRedAnno+: Annotation Exploitation in Self-Explanatory Lung Nodule Diagnosis
Paper and Code
Nov 07, 2022
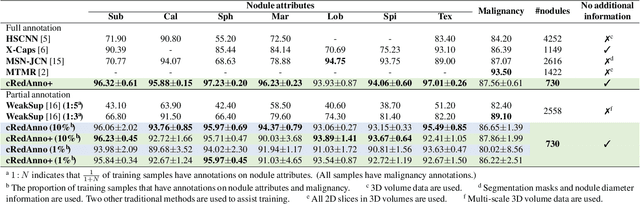
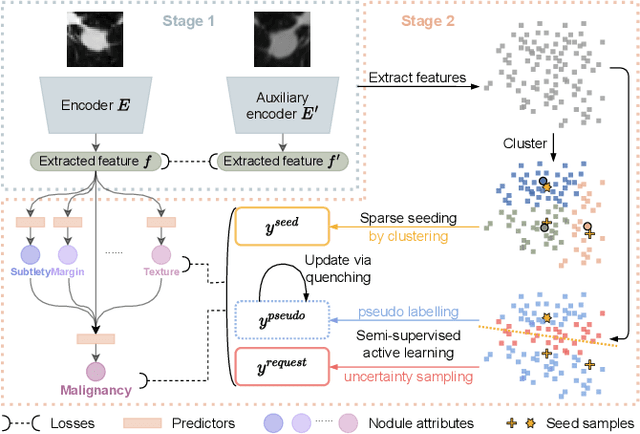
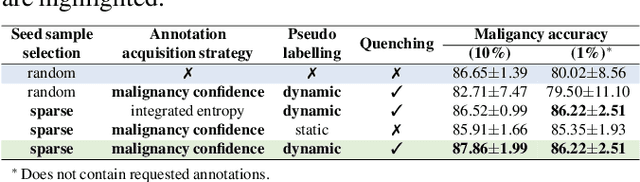
Recently, attempts have been made to reduce annotation requirements in feature-based self-explanatory models for lung nodule diagnosis. As a representative, cRedAnno achieves competitive performance with considerably reduced annotation needs by introducing self-supervised contrastive learning to do unsupervised feature extraction. However, it exhibits unstable performance under scarce annotation conditions. To improve the accuracy and robustness of cRedAnno, we propose an annotation exploitation mechanism by conducting semi-supervised active learning with sparse seeding and training quenching in the learned semantically meaningful reasoning space to jointly utilise the extracted features, annotations, and unlabelled data. The proposed approach achieves comparable or even higher malignancy prediction accuracy with 10x fewer annotations, meanwhile showing better robustness and nodule attribute prediction accuracy under the condition of 1% annotations. Our complete code is open-source available: https://github.com/diku-dk/credanno.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge