Brian Leahy
Developmental Stage Classification of Embryos Using Two-Stream Neural Network with Linear-Chain Conditional Random Field
Jul 13, 2021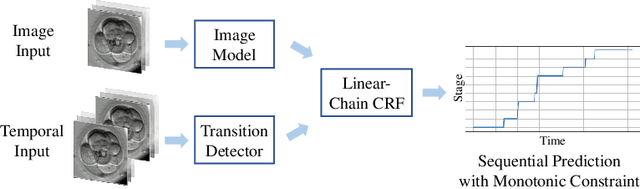
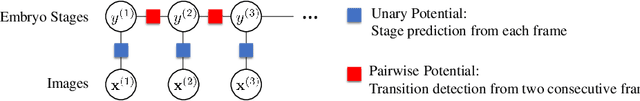
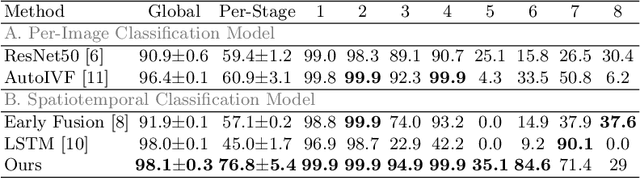

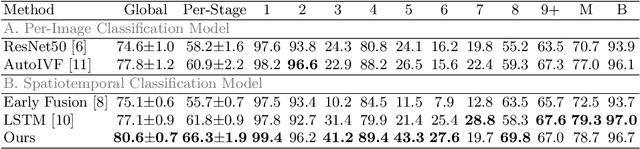
Abstract:The developmental process of embryos follows a monotonic order. An embryo can progressively cleave from one cell to multiple cells and finally transform to morula and blastocyst. For time-lapse videos of embryos, most existing developmental stage classification methods conduct per-frame predictions using an image frame at each time step. However, classification using only images suffers from overlapping between cells and imbalance between stages. Temporal information can be valuable in addressing this problem by capturing movements between neighboring frames. In this work, we propose a two-stream model for developmental stage classification. Unlike previous methods, our two-stream model accepts both temporal and image information. We develop a linear-chain conditional random field (CRF) on top of neural network features extracted from the temporal and image streams to make use of both modalities. The linear-chain CRF formulation enables tractable training of global sequential models over multiple frames while also making it possible to inject monotonic development order constraints into the learning process explicitly. We demonstrate our algorithm on two time-lapse embryo video datasets: i) mouse and ii) human embryo datasets. Our method achieves 98.1 % and 80.6 % for mouse and human embryo stage classification, respectively. Our approach will enable more profound clinical and biological studies and suggests a new direction for developmental stage classification by utilizing temporal information.
Learning Vector Quantized Shape Code for Amodal Blastomere Instance Segmentation
Dec 02, 2020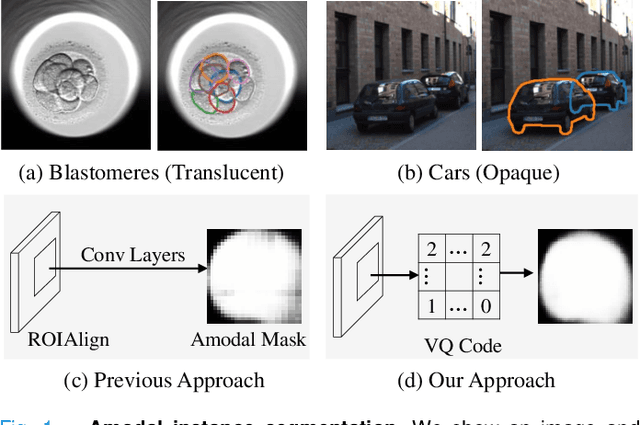



Abstract:Blastomere instance segmentation is important for analyzing embryos' abnormality. To measure the accurate shapes and sizes of blastomeres, their amodal segmentation is necessary. Amodal instance segmentation aims to recover the complete silhouette of an object even when the object is not fully visible. For each detected object, previous methods directly regress the target mask from input features. However, images of an object under different amounts of occlusion should have the same amodal mask output, which makes it harder to train the regression model. To alleviate the problem, we propose to classify input features into intermediate shape codes and recover complete object shapes from them. First, we pre-train the Vector Quantized Variational Autoencoder (VQ-VAE) model to learn these discrete shape codes from ground truth amodal masks. Then, we incorporate the VQ-VAE model into the amodal instance segmentation pipeline with an additional refinement module. We also detect an occlusion map to integrate occlusion information with a backbone feature. As such, our network faithfully detects bounding boxes of amodal objects. On an internal embryo cell image benchmark, the proposed method outperforms previous state-of-the-art methods. To show generalizability, we show segmentation results on the public KINS natural image benchmark. To examine the learned shape codes and model design choices, we perform ablation studies on a synthetic dataset of simple overlaid shapes. Our method would enable accurate measurement of blastomeres in in vitro fertilization (IVF) clinics, which potentially can increase IVF success rate.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge