Ardhendu Sekhar
Spatially-Aware Mixture of Experts with Log-Logistic Survival Modeling for Whole-Slide Images
Nov 17, 2025Abstract:Accurate survival prediction from histopathology whole-slide images (WSIs) remains challenging due to their gigapixel resolution, strong spatial heterogeneity, and complex survival distributions. We introduce a comprehensive computational pathology framework that addresses these limitations through four complementary innovations: (1) Quantile-Gated Patch Selection for dynamically identifying prognostically relevant regions, (2) Graph-Guided Clustering to group patches by spatial and morphological similarity, (3) Hierarchical Context Attention to model both local tissue interactions and global slide-level context, and (4) an Expert-Driven Mixture of Log-Logistics module that flexibly models complex survival distributions. Across large TCGA cohorts, our method achieves state-of-the-art performance, yielding time-dependent concordance indices of 0.644 on LUAD, 0.751 on KIRC, and 0.752 on BRCA, consistently outperforming both histology-only and multimodal baselines. The framework further provides improved calibration and interpretability, advancing the use of WSIs for personalized cancer prognosis.
Survival Modeling from Whole Slide Images via Patch-Level Graph Clustering and Mixture Density Experts
Jul 22, 2025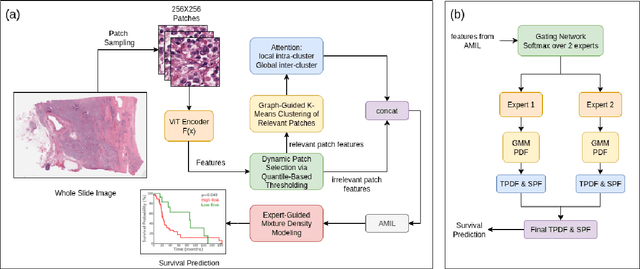
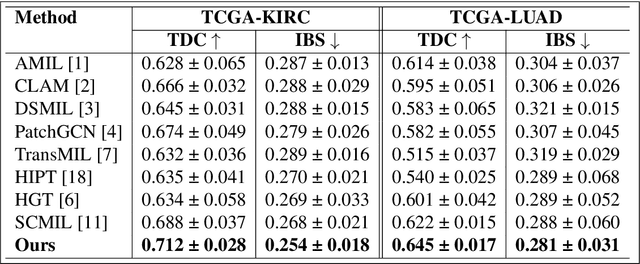
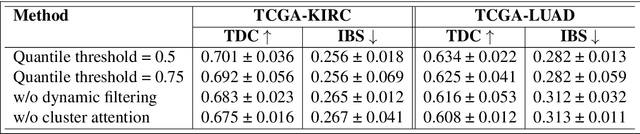
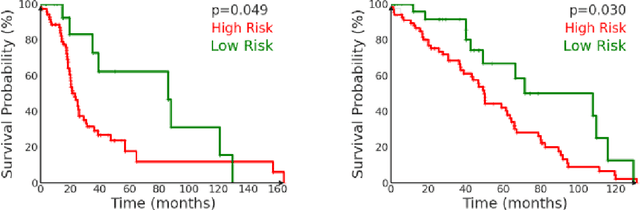
Abstract:We introduce a modular framework for predicting cancer-specific survival from whole slide pathology images (WSIs) that significantly improves upon the state-of-the-art accuracy. Our method integrating four key components. Firstly, to tackle large size of WSIs, we use dynamic patch selection via quantile-based thresholding for isolating prognostically informative tissue regions. Secondly, we use graph-guided k-means clustering to capture phenotype-level heterogeneity through spatial and morphological coherence. Thirdly, we use attention mechanisms that model both intra- and inter-cluster relationships to contextualize local features within global spatial relations between various types of tissue compartments. Finally, we use an expert-guided mixture density modeling for estimating complex survival distributions using Gaussian mixture models. The proposed model achieves a concordance index of $0.712 \pm 0.028$ and Brier score of $0.254 \pm 0.018$ on TCGA-KIRC (renal cancer), and a concordance index of $0.645 \pm 0.017$ and Brier score of $0.281 \pm 0.031$ on TCGA-LUAD (lung adenocarcinoma). These results are significantly better than the state-of-art and demonstrate predictive potential of the proposed method across diverse cancer types.
Predicting Genetic Mutations from Single-Cell Bone Marrow Images in Acute Myeloid Leukemia Using Noise-Robust Deep Learning Models
Jun 15, 2025Abstract:In this study, we propose a robust methodology for identification of myeloid blasts followed by prediction of genetic mutation in single-cell images of blasts, tackling challenges associated with label accuracy and data noise. We trained an initial binary classifier to distinguish between leukemic (blasts) and non-leukemic cells images, achieving 90 percent accuracy. To evaluate the models generalization, we applied this model to a separate large unlabeled dataset and validated the predictions with two haemato-pathologists, finding an approximate error rate of 20 percent in the leukemic and non-leukemic labels. Assuming this level of label noise, we further trained a four-class model on images predicted as blasts to classify specific mutations. The mutation labels were known for only a bag of cell images extracted from a single slide. Despite the tumor label noise, our mutation classification model achieved 85 percent accuracy across four mutation classes, demonstrating resilience to label inconsistencies. This study highlights the capability of machine learning models to work with noisy labels effectively while providing accurate, clinically relevant mutation predictions, which is promising for diagnostic applications in areas such as haemato-pathology.
Scalable Whole Slide Image Representation Using K-Mean Clustering and Fisher Vector Aggregation
Jan 21, 2025


Abstract:Whole slide images (WSIs) are high-resolution, gigapixel sized images that pose significant computational challenges for traditional machine learning models due to their size and heterogeneity.In this paper, we present a scalable and efficient methodology for WSI classification by leveraging patch-based feature extraction, clustering, and Fisher vector encoding. Initially, WSIs are divided into fixed size patches, and deep feature embeddings are extracted from each patch using a pre-trained convolutional neural network (CNN). These patch-level embeddings are subsequently clustered using K-means clustering, where each cluster aggregates semantically similar regions of the WSI. To effectively summarize each cluster, Fisher vector representations are computed by modeling the distribution of patch embeddings in each cluster as a parametric Gaussian mixture model (GMM). The Fisher vectors from each cluster are concatenated into a high-dimensional feature vector, creating a compact and informative representation of the entire WSI. This feature vector is then used by a classifier to predict the WSI's diagnostic label. Our method captures local and global tissue structures and yields robust performance for large-scale WSI classification, demonstrating superior accuracy and scalability compared to other approaches.
Cross-Domain Evaluation of Few-Shot Classification Models: Natural Images vs. Histopathological Images
Oct 11, 2024

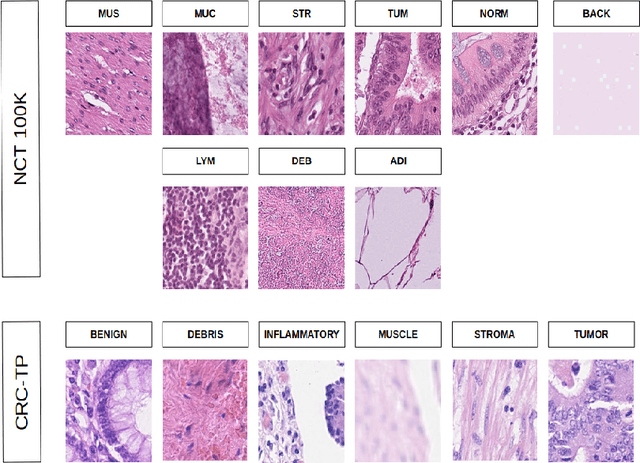

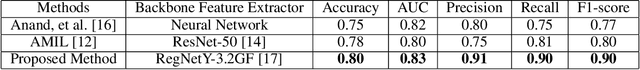
Abstract:In this study, we investigate the performance of few-shot classification models across different domains, specifically natural images and histopathological images. We first train several few-shot classification models on natural images and evaluate their performance on histopathological images. Subsequently, we train the same models on histopathological images and compare their performance. We incorporated four histopathology datasets and one natural images dataset and assessed performance across 5-way 1-shot, 5-way 5-shot, and 5-way 10-shot scenarios using a selection of state-of-the-art classification techniques. Our experimental results reveal insights into the transferability and generalization capabilities of few-shot classification models between diverse image domains. We analyze the strengths and limitations of these models in adapting to new domains and provide recommendations for optimizing their performance in cross-domain scenarios. This research contributes to advancing our understanding of few-shot learning in the context of image classification across diverse domains.
Few-Shot Histopathology Image Classification: Evaluating State-of-the-Art Methods and Unveiling Performance Insights
Aug 25, 2024



Abstract:This paper presents a study on few-shot classification in the context of histopathology images. While few-shot learning has been studied for natural image classification, its application to histopathology is relatively unexplored. Given the scarcity of labeled data in medical imaging and the inherent challenges posed by diverse tissue types and data preparation techniques, this research evaluates the performance of state-of-the-art few-shot learning methods for various scenarios on histology data. We have considered four histopathology datasets for few-shot histopathology image classification and have evaluated 5-way 1-shot, 5-way 5-shot and 5-way 10-shot scenarios with a set of state-of-the-art classification techniques. The best methods have surpassed an accuracy of 70%, 80% and 85% in the cases of 5-way 1-shot, 5-way 5-shot and 5-way 10-shot cases, respectively. We found that for histology images popular meta-learning approaches is at par with standard fine-tuning and regularization methods. Our experiments underscore the challenges of working with images from different domains and underscore the significance of unbiased and focused evaluations in advancing computer vision techniques for specialized domains, such as histology images.
HER2 and FISH Status Prediction in Breast Biopsy H&E-Stained Images Using Deep Learning
Aug 25, 2024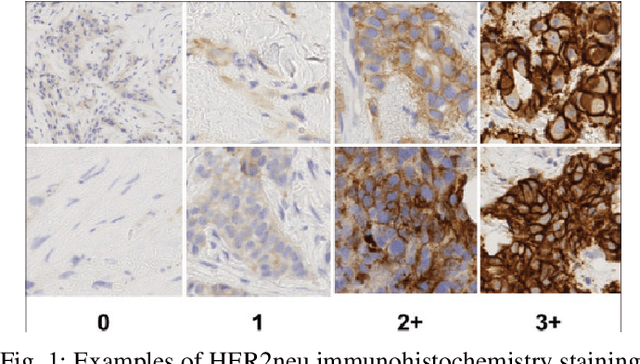
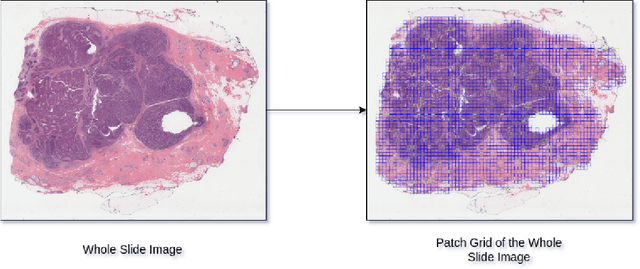

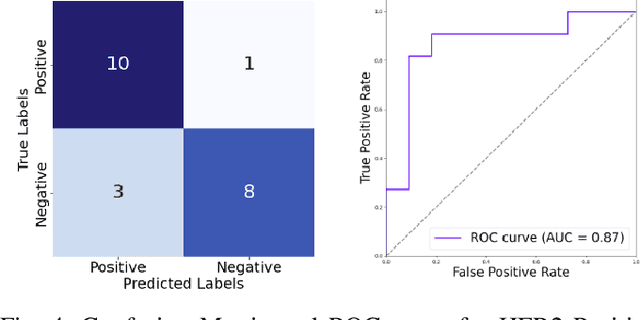
Abstract:The current standard for detecting human epidermal growth factor receptor 2 (HER2) status in breast cancer patients relies on HER2 amplification, identified through fluorescence in situ hybridization (FISH) or immunohistochemistry (IHC). However, hematoxylin and eosin (H\&E) tumor stains are more widely available, and accurately predicting HER2 status using H\&E could reduce costs and expedite treatment selection. Deep Learning algorithms for H&E have shown effectiveness in predicting various cancer features and clinical outcomes, including moderate success in HER2 status prediction. In this work, we employed a customized weak supervision classification technique combined with MoCo-v2 contrastive learning to predict HER2 status. We trained our pipeline on 182 publicly available H&E Whole Slide Images (WSIs) from The Cancer Genome Atlas (TCGA), for which annotations by the pathology team at Yale School of Medicine are publicly available. Our pipeline achieved an Area Under the Curve (AUC) of 0.85 across four different test folds. Additionally, we tested our model on 44 H&E slides from the TCGA-BRCA dataset, which had an HER2 score of 2+ and included corresponding HER2 status and FISH test results. These cases are considered equivocal for IHC, requiring an expensive FISH test on their IHC slides for disambiguation. Our pipeline demonstrated an AUC of 0.81 on these challenging H&E slides. Reducing the need for FISH test can have significant implications in cancer treatment equity for underserved populations.
IFSENet : Harnessing Sparse Iterations for Interactive Few-shot Segmentation Excellence
Mar 22, 2024Abstract:Training a computer vision system to segment a novel class typically requires collecting and painstakingly annotating lots of images with objects from that class. Few-shot segmentation techniques reduce the required number of images to learn to segment a new class, but careful annotations of object boundaries are still required. On the other hand, interactive segmentation techniques only focus on incrementally improving the segmentation of one object at a time (typically, using clicks given by an expert) in a class-agnostic manner. We combine the two concepts to drastically reduce the effort required to train segmentation models for novel classes. Instead of trivially feeding interactive segmentation masks as ground truth to a few-shot segmentation model, we propose IFSENet, which can accept sparse supervision on a single or few support images in the form of clicks to generate masks on support (training, at least clicked upon once) as well as query (test, never clicked upon) images. To trade-off effort for accuracy flexibly, the number of images and clicks can be incrementally added to the support set to further improve the segmentation of support as well as query images. The proposed model approaches the accuracy of previous state-of-the-art few-shot segmentation models with considerably lower annotation effort (clicks instead of maps), when tested on Pascal and SBD datasets on query images. It also works well as an interactive segmentation method on support images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge