Alexey V. Dimov
Synthetic Generation and Latent Projection Denoising of Rim Lesions in Multiple Sclerosis
May 29, 2025



Abstract:Quantitative susceptibility maps from magnetic resonance images can provide both prognostic and diagnostic information in multiple sclerosis, a neurodegenerative disease characterized by the formation of lesions in white matter brain tissue. In particular, susceptibility maps provide adequate contrast to distinguish between "rim" lesions, surrounded by deposited paramagnetic iron, and "non-rim" lesion types. These paramagnetic rim lesions (PRLs) are an emerging biomarker in multiple sclerosis. Much effort has been devoted to both detection and segmentation of such lesions to monitor longitudinal change. As paramagnetic rim lesions are rare, addressing this problem requires confronting the class imbalance between rim and non-rim lesions. We produce synthetic quantitative susceptibility maps of paramagnetic rim lesions and show that inclusion of such synthetic data improves classifier performance and provide a multi-channel extension to generate accompanying contrasts and probabilistic segmentation maps. We exploit the projection capability of our trained generative network to demonstrate a novel denoising approach that allows us to train on ambiguous rim cases and substantially increase the minority class. We show that both synthetic lesion synthesis and our proposed rim lesion label denoising method best approximate the unseen rim lesion distribution and improve detection in a clinically interpretable manner. We release our code and generated data at https://github.com/agr78/PRLx-GAN upon publication.
Implicit neural representation for free-breathing MR fingerprinting (INR-MRF): co-registered 3D whole-liver water T1, water T2, proton density fat fraction, and R2* mapping
Oct 19, 2024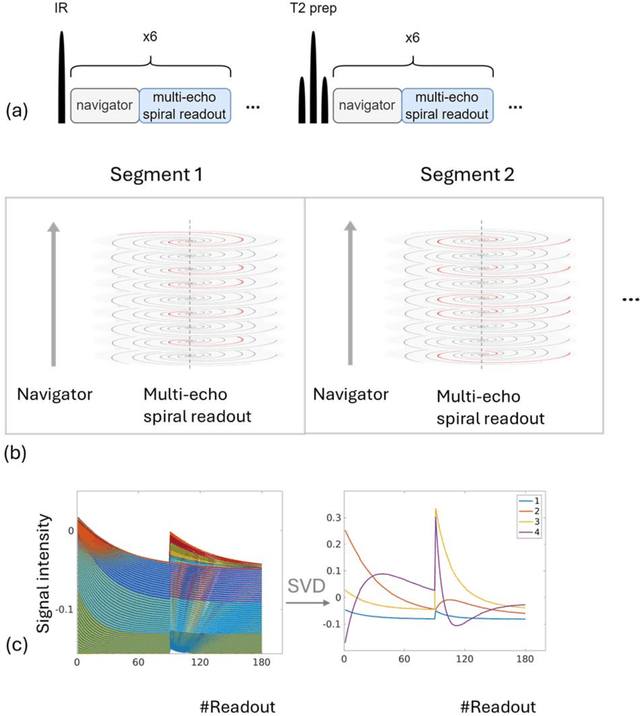
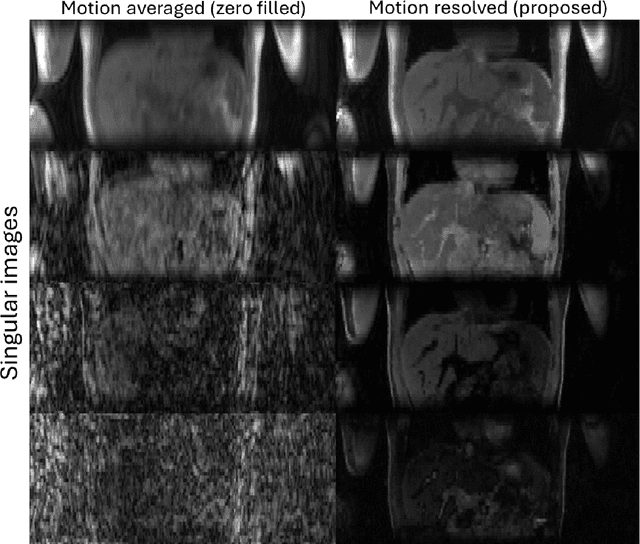

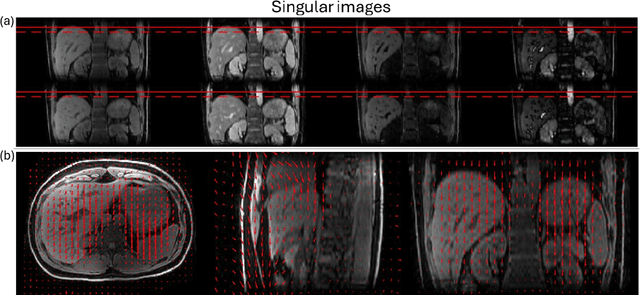

Abstract:Purpose: To develop an MRI technique for free-breathing 3D whole-liver quantification of water T1, water T2, proton density fat fraction (PDFF), R2*. Methods: An Eight-echo spoiled gradient echo pulse sequence with spiral readout was developed by interleaving inversion recovery and T2 magnetization preparation. We propose a neural network based on a 4D and a 3D implicit neural representation (INR) which simultaneously learns the motion deformation fields and the static reference frame MRI subspace images respectively. Water and fat singular images were separated during network training, with no need of performing retrospective water-fat separation. T1, T2, R2* and proton density fat fraction (PDFF) produced by the proposed method were validated in vivo on 10 healthy subjects, using quantitative maps generated from conventional scans as reference. Results: Our results showed minimal bias and narrow 95% limits of agreement on T1, T2, R2* and PDFF values in the liver compared to conventional breath-holding scans. Conclusions: INR-MRF enabled co-registered 3D whole liver T1, T2, R2* and PDFF mapping in a single free-breathing scan.
MRI quantification of liver fibrosis using diamagnetic susceptibility: An ex-vivo feasibility study
Oct 04, 2024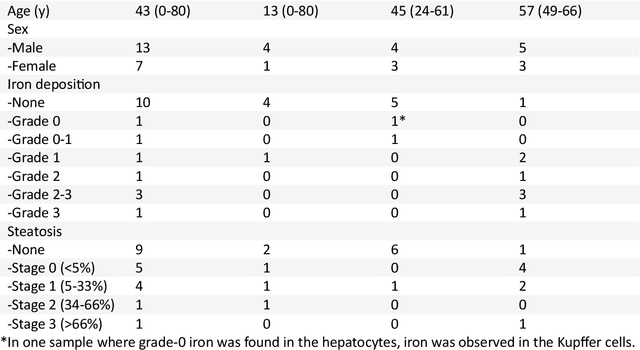
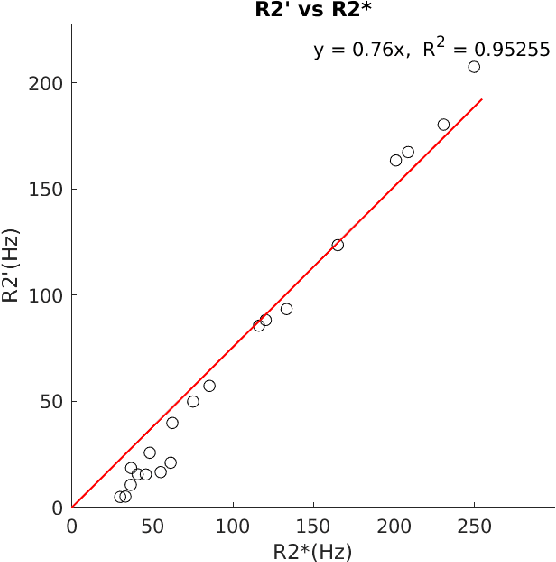
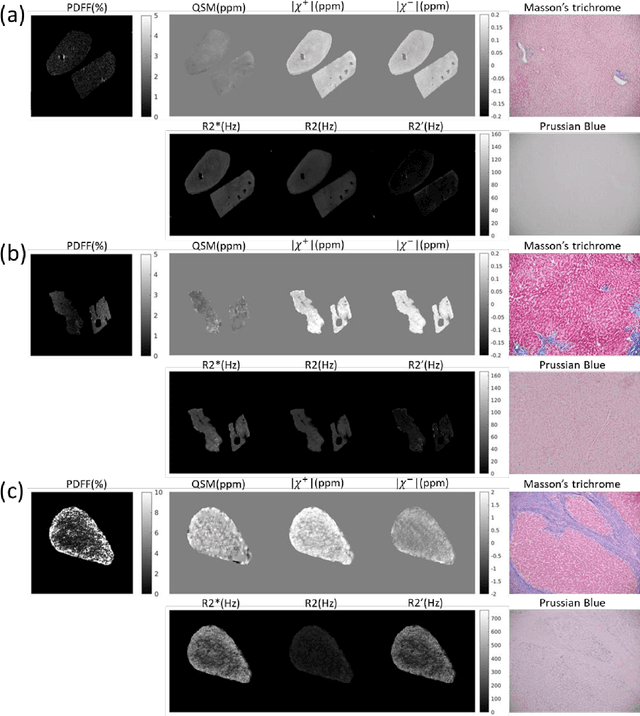
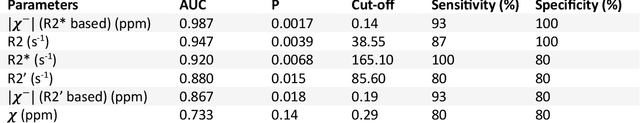
Abstract:In chronic liver disease, liver fibrosis develops as excessive deposition of extracellular matrix macromolecules, predominantly collagens, progressively form fibrous scars that disrupt the hepatic architecture, and fibrosis, iron, and fat are interrelated. Fibrosis is the best predictor of morbidity and mortality in chronic liver disease but liver biopsy, the reference method for diagnosis and staging, is invasive and limited by sampling and interobserver variability and risks of complications. The overall objective of this study was to develop a new non-invasive method to quantify fibrosis using diamagnetic susceptibility sources with histology validation in ex vivo liver explants.
Maximum Spherical Mean Value (mSMV) Filtering for Whole Brain Quantitative Susceptibility Mapping
Apr 22, 2023



Abstract:To develop a tissue field filtering algorithm, called maximum Spherical Mean Value (mSMV), for reducing shadow artifacts in quantitative susceptibility mapping (QSM) of the brain without requiring brain tissue erosion. Residual background field is a major source of shadow artifacts in QSM. The mSMV algorithm filters large field values near the border, where the maximum value of the harmonic background field is located. The effectiveness of mSMV for artifact removal was evaluated by comparing with existing QSM algorithms in a simulated numerical brain, 11 healthy volunteers, by assessing image quality in routine clinical patient study $(n=43)$, and by measuring lesion susceptibility values in multiple sclerosis patients $(n=50)$, a total of $n=93$ patients. Numerical simulation showed that mSMV reduces shadow artifacts and improves QSM accuracy. Better shadow reduction, as demonstrated by lower QSM variation in the gray matter and higher QSM image quality score, was also observed in healthy subjects and in patients with hemorrhages, stroke and multiple sclerosis. The mSMV algorithm allows reconstruction of QSMs that are equivalent to those obtained using SMV-filtered dipole inversion without eroding the volume of interest.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge