Akshay Smit
Effect of Radiology Report Labeler Quality on Deep Learning Models for Chest X-Ray Interpretation
Apr 01, 2021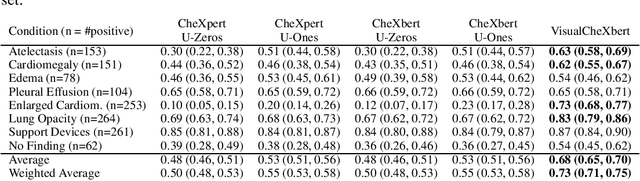

Abstract:Although deep learning models for chest X-ray interpretation are commonly trained on labels generated by automatic radiology report labelers, the impact of improvements in report labeling on the performance of chest X-ray classification models has not been systematically investigated. We first compare the CheXpert, CheXbert, and VisualCheXbert labelers on the task of extracting accurate chest X-ray image labels from radiology reports, reporting that the VisualCheXbert labeler outperforms the CheXpert and CheXbert labelers. Next, after training image classification models using labels generated from the different radiology report labelers on one of the largest datasets of chest X-rays, we show that an image classification model trained on labels from the VisualCheXbert labeler outperforms image classification models trained on labels from the CheXpert and CheXbert labelers. Our work suggests that recent improvements in radiology report labeling can translate to the development of higher performing chest X-ray classification models.
MedSelect: Selective Labeling for Medical Image Classification Combining Meta-Learning with Deep Reinforcement Learning
Mar 26, 2021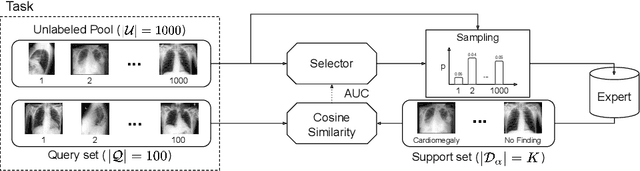


Abstract:We propose a selective learning method using meta-learning and deep reinforcement learning for medical image interpretation in the setting of limited labeling resources. Our method, MedSelect, consists of a trainable deep learning selector that uses image embeddings obtained from contrastive pretraining for determining which images to label, and a non-parametric selector that uses cosine similarity to classify unseen images. We demonstrate that MedSelect learns an effective selection strategy outperforming baseline selection strategies across seen and unseen medical conditions for chest X-ray interpretation. We also perform an analysis of the selections performed by MedSelect comparing the distribution of latent embeddings and clinical features, and find significant differences compared to the strongest performing baseline. We believe that our method may be broadly applicable across medical imaging settings where labels are expensive to acquire.
VisualCheXbert: Addressing the Discrepancy Between Radiology Report Labels and Image Labels
Mar 15, 2021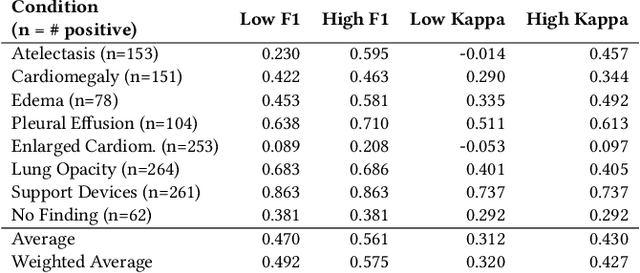
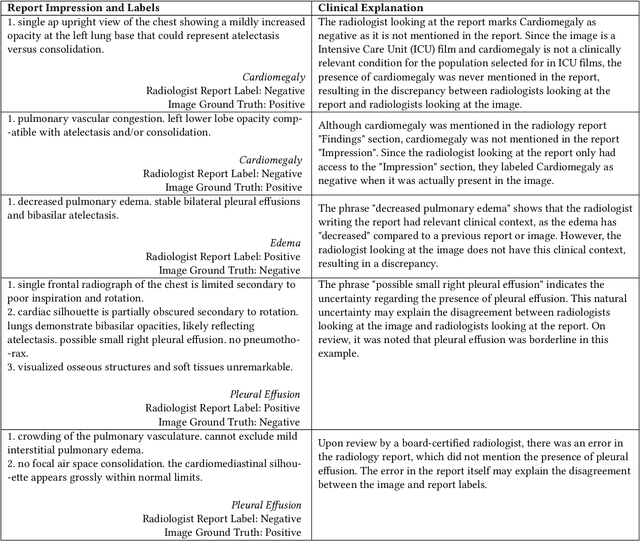

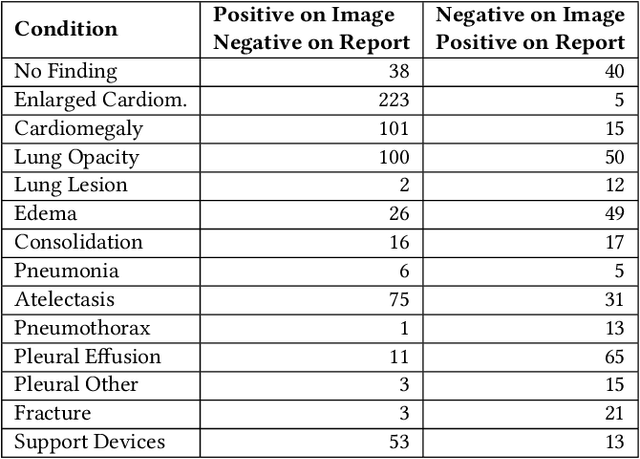
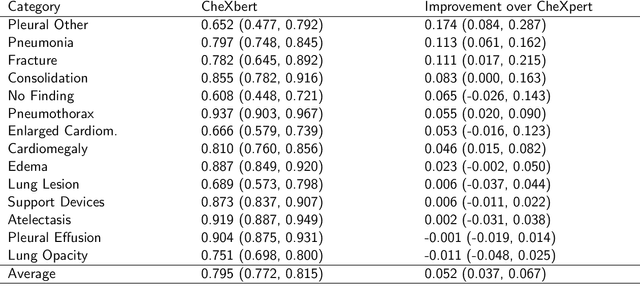
Abstract:Automatic extraction of medical conditions from free-text radiology reports is critical for supervising computer vision models to interpret medical images. In this work, we show that radiologists labeling reports significantly disagree with radiologists labeling corresponding chest X-ray images, which reduces the quality of report labels as proxies for image labels. We develop and evaluate methods to produce labels from radiology reports that have better agreement with radiologists labeling images. Our best performing method, called VisualCheXbert, uses a biomedically-pretrained BERT model to directly map from a radiology report to the image labels, with a supervisory signal determined by a computer vision model trained to detect medical conditions from chest X-ray images. We find that VisualCheXbert outperforms an approach using an existing radiology report labeler by an average F1 score of 0.14 (95% CI 0.12, 0.17). We also find that VisualCheXbert better agrees with radiologists labeling chest X-ray images than do radiologists labeling the corresponding radiology reports by an average F1 score across several medical conditions of between 0.12 (95% CI 0.09, 0.15) and 0.21 (95% CI 0.18, 0.24).
DLBCL-Morph: Morphological features computed using deep learning for an annotated digital DLBCL image set
Sep 24, 2020
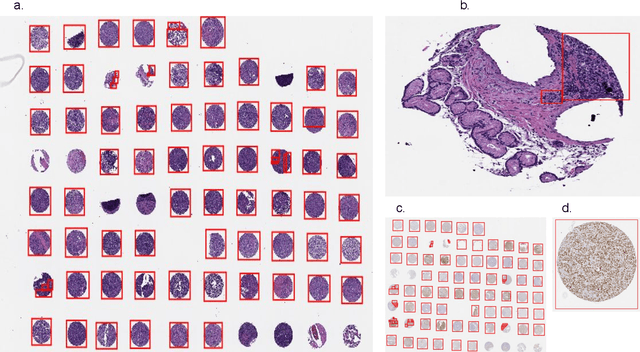

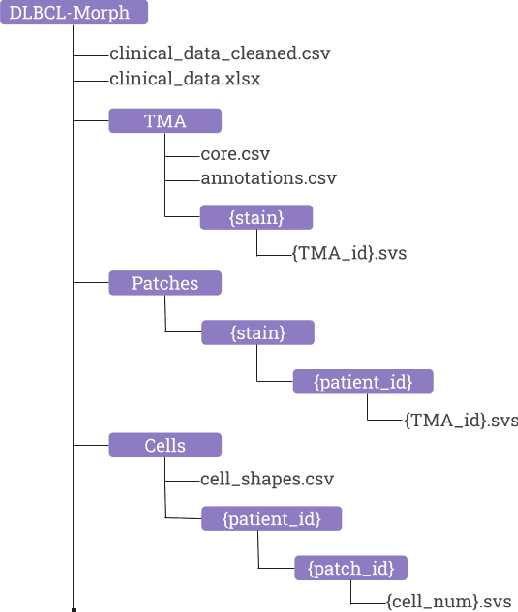
Abstract:Diffuse Large B-Cell Lymphoma (DLBCL) is the most common non-Hodgkin lymphoma. Though histologically DLBCL shows varying morphologies, no morphologic features have been consistently demonstrated to correlate with prognosis. We present a morphologic analysis of histology sections from 209 DLBCL cases with associated clinical and cytogenetic data. Duplicate tissue core sections were arranged in tissue microarrays (TMAs), and replicate sections were stained with H&E and immunohistochemical stains for CD10, BCL6, MUM1, BCL2, and MYC. The TMAs are accompanied by pathologist-annotated regions-of-interest (ROIs) that identify areas of tissue representative of DLBCL. We used a deep learning model to segment all tumor nuclei in the ROIs, and computed several geometric features for each segmented nucleus. We fit a Cox proportional hazards model to demonstrate the utility of these geometric features in predicting survival outcome, and found that it achieved a C-index (95% CI) of 0.635 (0.574,0.691). Our finding suggests that geometric features computed from tumor nuclei are of prognostic importance, and should be validated in prospective studies.
CheXbert: Combining Automatic Labelers and Expert Annotations for Accurate Radiology Report Labeling Using BERT
Apr 30, 2020

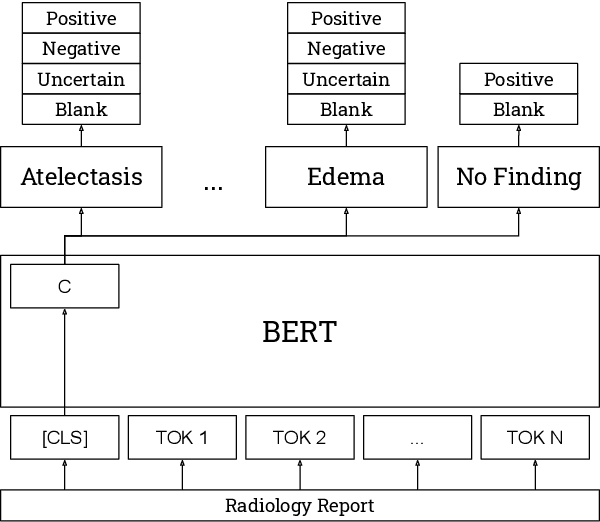
Abstract:The extraction of labels from radiology text reports enables large-scale training of medical imaging models. Existing approaches to report labeling typically rely either on sophisticated feature engineering based on medical domain knowledge or manual annotations by experts. In this work, we introduce a BERT-based approach to medical image report labeling that exploits both the scale of available rule-based systems and the quality of expert annotations. We demonstrate superior performance of a biomedically pretrained BERT model first trained on annotations of a rule-based labeler and then finetuned on a small set of expert annotations augmented with automated backtranslation. We find that our final model, CheXbert, is able to outperform the previous best rules-based labeler with statistical significance, setting a new SOTA for report labeling on one of the largest datasets of chest x-rays.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge