Reliable Liver Fibrosis Assessment from Ultrasound using Global Hetero-Image Fusion and View-Specific Parameterization
Paper and Code
Aug 07, 2020
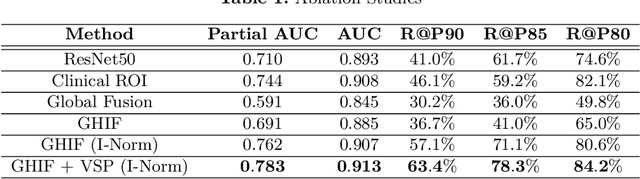
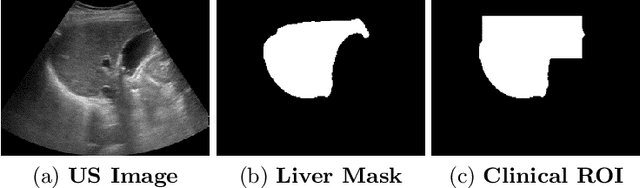
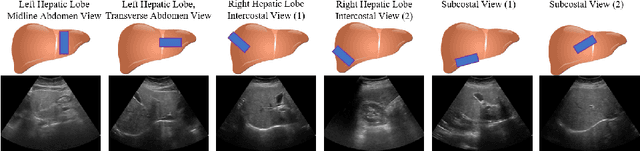
Ultrasound (US) is a critical modality for diagnosing liver fibrosis. Unfortunately, assessment is very subjective, motivating automated approaches. We introduce a principled deep convolutional neural network (CNN) workflow that incorporates several innovations. First, to avoid overfitting on non-relevant image features, we force the network to focus on a clinical region of interest (ROI), encompassing the liver parenchyma and upper border. Second, we introduce global heteroimage fusion (GHIF), which allows the CNN to fuse features from any arbitrary number of images in a study, increasing its versatility and flexibility. Finally, we use 'style'-based view-specific parameterization (VSP) to tailor the CNN processing for different viewpoints of the liver, while keeping the majority of parameters the same across views. Experiments on a dataset of 610 patient studies (6979 images) demonstrate that our pipeline can contribute roughly 7% and 22% improvements in partial area under the curve and recall at 90% precision, respectively, over conventional classifiers, validating our approach to this crucial problem.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge