ProtIR: Iterative Refinement between Retrievers and Predictors for Protein Function Annotation
Paper and Code
Feb 10, 2024
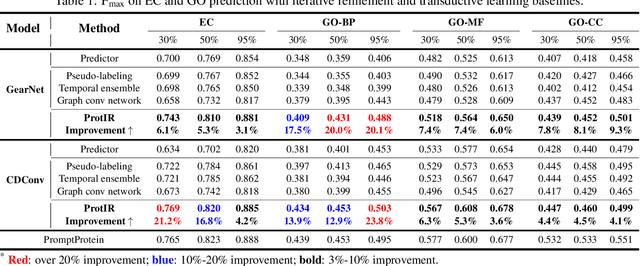
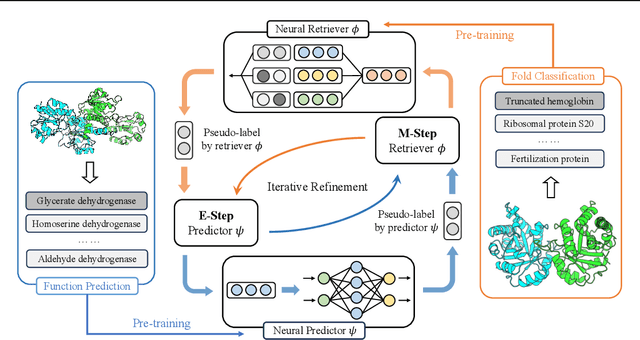

Protein function annotation is an important yet challenging task in biology. Recent deep learning advancements show significant potential for accurate function prediction by learning from protein sequences and structures. Nevertheless, these predictor-based methods often overlook the modeling of protein similarity, an idea commonly employed in traditional approaches using sequence or structure retrieval tools. To fill this gap, we first study the effect of inter-protein similarity modeling by benchmarking retriever-based methods against predictors on protein function annotation tasks. Our results show that retrievers can match or outperform predictors without large-scale pre-training. Building on these insights, we introduce a novel variational pseudo-likelihood framework, ProtIR, designed to improve function predictors by incorporating inter-protein similarity modeling. This framework iteratively refines knowledge between a function predictor and retriever, thereby combining the strengths of both predictors and retrievers. ProtIR showcases around 10% improvement over vanilla predictor-based methods. Besides, it achieves performance on par with protein language model-based methods, yet without the need for massive pre-training, highlighting the efficacy of our framework. Code will be released upon acceptance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge