MolNexTR: A Generalized Deep Learning Model for Molecular Image Recognition
Paper and Code
Mar 08, 2024

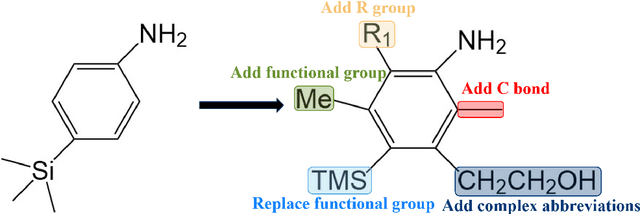
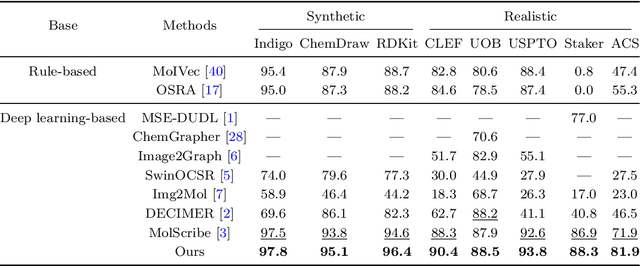
In the field of chemical structure recognition, the task of converting molecular images into graph structures and SMILES string stands as a significant challenge, primarily due to the varied drawing styles and conventions prevalent in chemical literature. To bridge this gap, we proposed MolNexTR, a novel image-to-graph deep learning model that collaborates to fuse the strengths of ConvNext, a powerful Convolutional Neural Network variant, and Vision-TRansformer. This integration facilitates a more nuanced extraction of both local and global features from molecular images. MolNexTR can predict atoms and bonds simultaneously and understand their layout rules. It also excels at flexibly integrating symbolic chemistry principles to discern chirality and decipher abbreviated structures. We further incorporate a series of advanced algorithms, including improved data augmentation module, image contamination module, and a post-processing module to get the final SMILES output. These modules synergistically enhance the model's robustness against the diverse styles of molecular imagery found in real literature. In our test sets, MolNexTR has demonstrated superior performance, achieving an accuracy rate of 81-97%, marking a significant advancement in the domain of molecular structure recognition. Scientific contribution: MolNexTR is a novel image-to-graph model that incorporates a unique dual-stream encoder to extract complex molecular image features, and combines chemical rules to predict atoms and bonds while understanding atom and bond layout rules. In addition, it employs a series of novel augmentation algorithms to significantly enhance the robustness and performance of the model.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge