Meta Mask Correction for Nuclei Segmentation in Histopathological Image
Paper and Code
Nov 24, 2021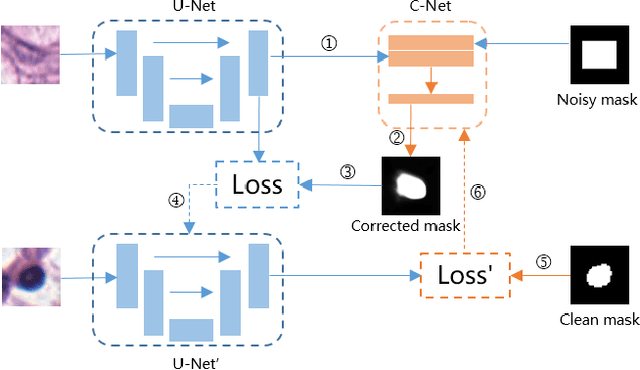
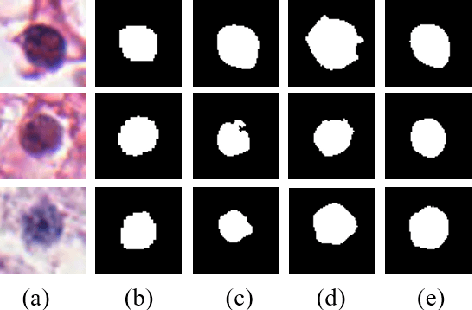

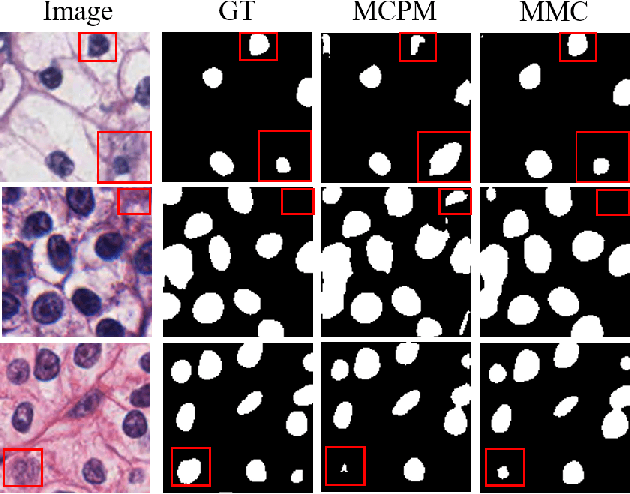
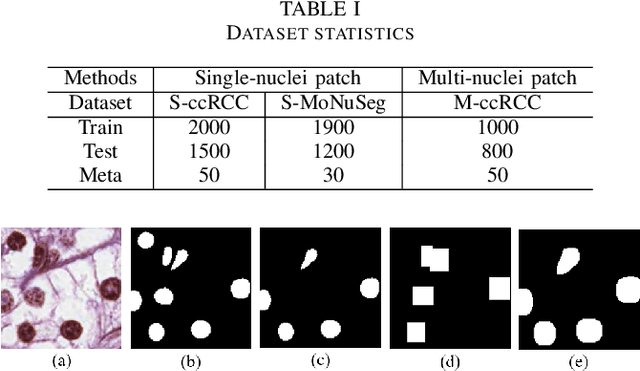
Nuclei segmentation is a fundamental task in digital pathology analysis and can be automated by deep learning-based methods. However, the development of such an automated method requires a large amount of data with precisely annotated masks which is hard to obtain. Training with weakly labeled data is a popular solution for reducing the workload of annotation. In this paper, we propose a novel meta-learning-based nuclei segmentation method which follows the label correction paradigm to leverage data with noisy masks. Specifically, we design a fully conventional meta-model that can correct noisy masks using a small amount of clean meta-data. Then the corrected masks can be used to supervise the training of the segmentation model. Meanwhile, a bi-level optimization method is adopted to alternately update the parameters of the main segmentation model and the meta-model in an end-to-end way. Extensive experimental results on two nuclear segmentation datasets show that our method achieves the state-of-the-art result. It even achieves comparable performance with the model training on supervised data in some noisy settings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge