A Novel Framework Integrating AI Model and Enzymological Experiments Promotes Identification of SARS-CoV-2 3CL Protease Inhibitors and Activity-based Probe
Paper and Code
May 29, 2021
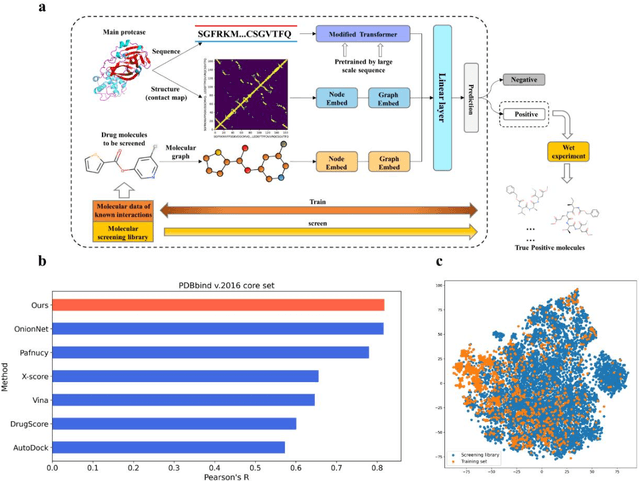
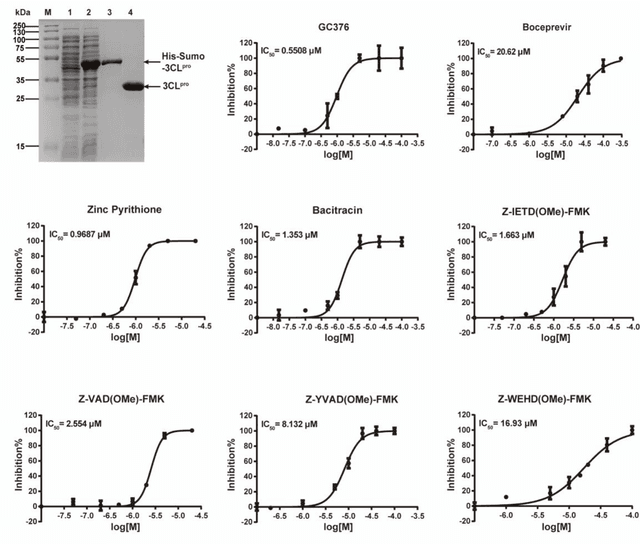
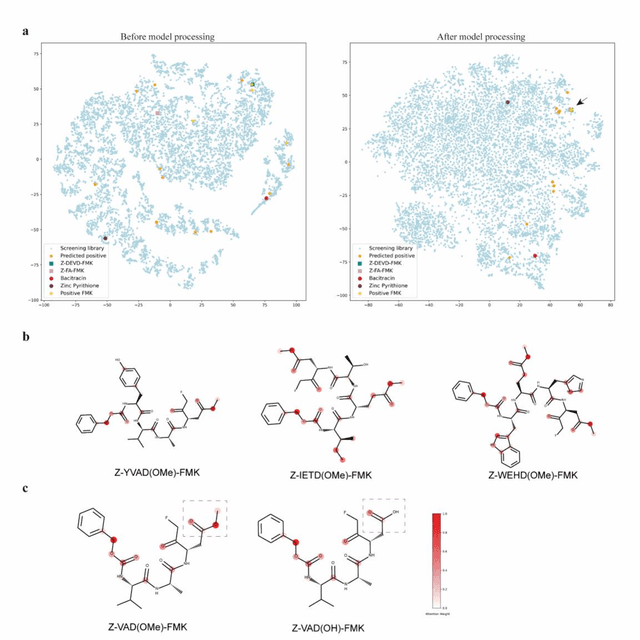
The identification of protein-ligand interaction plays a key role in biochemical research and drug discovery. Although deep learning has recently shown great promise in discovering new drugs, there remains a gap between deep learning-based and experimental approaches. Here we propose a novel framework, named AIMEE, integrating AI Model and Enzymology Experiments, to identify inhibitors against 3CL protease of SARS-CoV-2, which has taken a significant toll on people across the globe. From a bioactive chemical library, we have conducted two rounds of experiments and identified six novel inhibitors with a hit rate of 29.41%, and four of them showed an IC50 value less than 3 {\mu}M. Moreover, we explored the interpretability of the central model in AIMEE, mapping the deep learning extracted features to domain knowledge of chemical properties. Based on this knowledge, a commercially available compound was selected and proven to be an activity-based probe of 3CLpro. This work highlights the great potential of combining deep learning models and biochemical experiments for intelligent iteration and expanding the boundaries of drug discovery.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge