Ziya Ata Yazıcı
A Survey on Class-Agnostic Counting: Advancements from Reference-Based to Open-World Text-Guided Approaches
Jan 31, 2025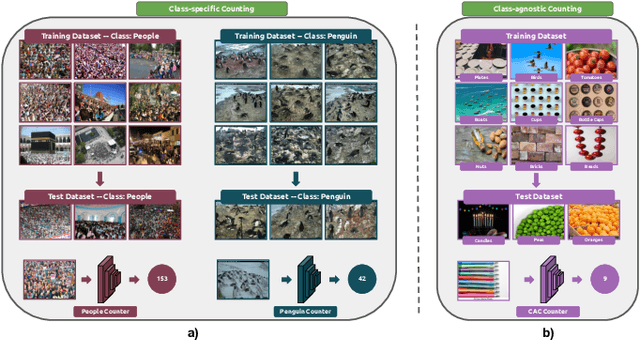
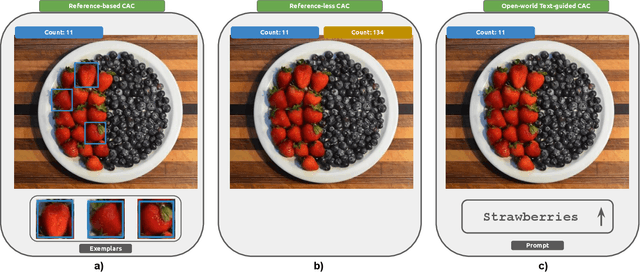

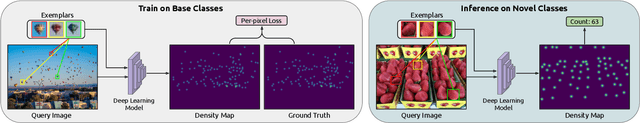
Abstract:Object counting has recently shifted towards class-agnostic counting (CAC), which addresses the challenge of counting objects across arbitrary categories, tackling a critical need in versatile counting systems. While humans effortlessly identify and count objects from diverse categories without prior knowledge, most counting methods remain restricted to enumerating instances of known classes, requiring extensive labeled datasets for training, and struggling under open-vocabulary settings. Conversely, CAC aims to count objects belonging to classes never seen during training, typically operating in a few-shot setting. In this paper, for the first time, we review advancements in CAC methodologies, categorizing them into three paradigms based on how target object classes can be specified: reference-based, reference-less, and open-world text-guided. Reference-based approaches have set performance benchmarks using exemplar-guided mechanisms. Reference-less methods eliminate exemplar dependency by leveraging inherent image patterns. Finally, open-world text-guided methods utilize vision-language models, enabling object class descriptions through textual prompts, representing a flexible and appealing solution. We analyze state-of-the-art techniques and we report their results on existing gold standard benchmarks, comparing their performance and identifying and discussing their strengths and limitations. Persistent challenges -- such as annotation dependency, scalability, and generalization -- are discussed, alongside future directions. We believe this survey serves as a valuable resource for researchers to understand the progressive developments and contributions over time and the current state-of-the-art of CAC, suggesting insights for future directions and challenges to be addressed.
GLIMS: Attention-Guided Lightweight Multi-Scale Hybrid Network for Volumetric Semantic Segmentation
Apr 27, 2024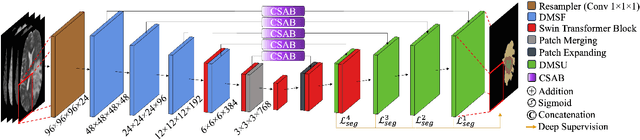



Abstract:Convolutional Neural Networks (CNNs) have become widely adopted for medical image segmentation tasks, demonstrating promising performance. However, the inherent inductive biases in convolutional architectures limit their ability to model long-range dependencies and spatial correlations. While recent transformer-based architectures address these limitations by leveraging self-attention mechanisms to encode long-range dependencies and learn expressive representations, they often struggle to extract low-level features and are highly dependent on data availability. This motivated us for the development of GLIMS, a data-efficient attention-guided hybrid volumetric segmentation network. GLIMS utilizes Dilated Feature Aggregator Convolutional Blocks (DACB) to capture local-global feature correlations efficiently. Furthermore, the incorporated Swin Transformer-based bottleneck bridges the local and global features to improve the robustness of the model. Additionally, GLIMS employs an attention-guided segmentation approach through Channel and Spatial-Wise Attention Blocks (CSAB) to localize expressive features for fine-grained border segmentation. Quantitative and qualitative results on glioblastoma and multi-organ CT segmentation tasks demonstrate GLIMS' effectiveness in terms of complexity and accuracy. GLIMS demonstrated outstanding performance on BraTS2021 and BTCV datasets, surpassing the performance of Swin UNETR. Notably, GLIMS achieved this high performance with a significantly reduced number of trainable parameters. Specifically, GLIMS has 47.16M trainable parameters and 72.30G FLOPs, while Swin UNETR has 61.98M trainable parameters and 394.84G FLOPs. The code is publicly available on https://github.com/yaziciz/GLIMS.
Attention-Enhanced Hybrid Feature Aggregation Network for 3D Brain Tumor Segmentation
Mar 15, 2024Abstract:Glioblastoma is a highly aggressive and malignant brain tumor type that requires early diagnosis and prompt intervention. Due to its heterogeneity in appearance, developing automated detection approaches is challenging. To address this challenge, Artificial Intelligence (AI)-driven approaches in healthcare have generated interest in efficiently diagnosing and evaluating brain tumors. The Brain Tumor Segmentation Challenge (BraTS) is a platform for developing and assessing automated techniques for tumor analysis using high-quality, clinically acquired MRI data. In our approach, we utilized a multi-scale, attention-guided and hybrid U-Net-shaped model -- GLIMS -- to perform 3D brain tumor segmentation in three regions: Enhancing Tumor (ET), Tumor Core (TC), and Whole Tumor (WT). The multi-scale feature extraction provides better contextual feature aggregation in high resolutions and the Swin Transformer blocks improve the global feature extraction at deeper levels of the model. The segmentation mask generation in the decoder branch is guided by the attention-refined features gathered from the encoder branch to enhance the important attributes. Moreover, hierarchical supervision is used to train the model efficiently. Our model's performance on the validation set resulted in 92.19, 87.75, and 83.18 Dice Scores and 89.09, 84.67, and 82.15 Lesion-wise Dice Scores in WT, TC, and ET, respectively. The code is publicly available at https://github.com/yaziciz/GLIMS.
In-Bed Pose Estimation: A Review
Feb 01, 2024Abstract:Human pose estimation, the process of identifying joint positions in a person's body from images or videos, represents a widely utilized technology across diverse fields, including healthcare. One such healthcare application involves in-bed pose estimation, where the body pose of an individual lying under a blanket is analyzed. This task, for instance, can be used to monitor a person's sleep behavior and detect symptoms early for potential disease diagnosis in homes and hospitals. Several studies have utilized unimodal and multimodal methods to estimate in-bed human poses. The unimodal studies generally employ RGB images, whereas the multimodal studies use modalities including RGB, long-wavelength infrared, pressure map, and depth map. Multimodal studies have the advantage of using modalities in addition to RGB that might capture information useful to cope with occlusions. Moreover, some multimodal studies exclude RGB and, this way, better suit privacy preservation. To expedite advancements in this domain, we conduct a review of existing datasets and approaches. Our objectives are to show the limitations of the previous studies, current challenges, and provide insights for future works on the in-bed human pose estimation field.
VIDI: A Video Dataset of Incidents
May 26, 2022
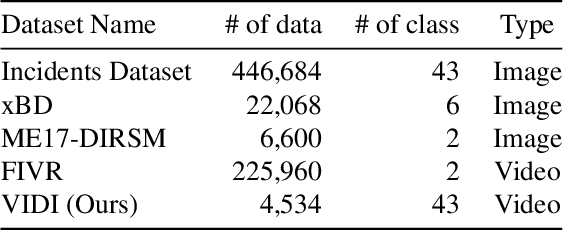
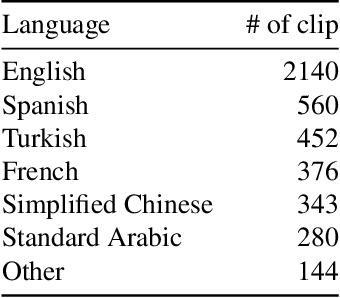

Abstract:Automatic detection of natural disasters and incidents has become more important as a tool for fast response. There have been many studies to detect incidents using still images and text. However, the number of approaches that exploit temporal information is rather limited. One of the main reasons for this is that a diverse video dataset with various incident types does not exist. To address this need, in this paper we present a video dataset, Video Dataset of Incidents, VIDI, that contains 4,534 video clips corresponding to 43 incident categories. Each incident class has around 100 videos with a duration of ten seconds on average. To increase diversity, the videos have been searched in several languages. To assess the performance of the recent state-of-the-art approaches, Vision Transformer and TimeSformer, as well as to explore the contribution of video-based information for incident classification, we performed benchmark experiments on the VIDI and Incidents Dataset. We have shown that the recent methods improve the incident classification accuracy. We have found that employing video data is very beneficial for the task. By using the video data, the top-1 accuracy is increased to 76.56% from 67.37%, which was obtained using a single frame. VIDI will be made publicly available. Additional materials can be found at the following link: https://github.com/vididataset/VIDI.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge