GLIMS: Attention-Guided Lightweight Multi-Scale Hybrid Network for Volumetric Semantic Segmentation
Paper and Code
Apr 27, 2024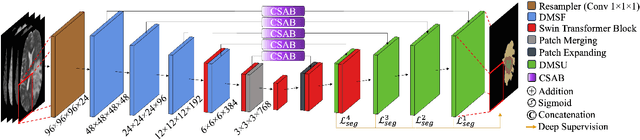



Convolutional Neural Networks (CNNs) have become widely adopted for medical image segmentation tasks, demonstrating promising performance. However, the inherent inductive biases in convolutional architectures limit their ability to model long-range dependencies and spatial correlations. While recent transformer-based architectures address these limitations by leveraging self-attention mechanisms to encode long-range dependencies and learn expressive representations, they often struggle to extract low-level features and are highly dependent on data availability. This motivated us for the development of GLIMS, a data-efficient attention-guided hybrid volumetric segmentation network. GLIMS utilizes Dilated Feature Aggregator Convolutional Blocks (DACB) to capture local-global feature correlations efficiently. Furthermore, the incorporated Swin Transformer-based bottleneck bridges the local and global features to improve the robustness of the model. Additionally, GLIMS employs an attention-guided segmentation approach through Channel and Spatial-Wise Attention Blocks (CSAB) to localize expressive features for fine-grained border segmentation. Quantitative and qualitative results on glioblastoma and multi-organ CT segmentation tasks demonstrate GLIMS' effectiveness in terms of complexity and accuracy. GLIMS demonstrated outstanding performance on BraTS2021 and BTCV datasets, surpassing the performance of Swin UNETR. Notably, GLIMS achieved this high performance with a significantly reduced number of trainable parameters. Specifically, GLIMS has 47.16M trainable parameters and 72.30G FLOPs, while Swin UNETR has 61.98M trainable parameters and 394.84G FLOPs. The code is publicly available on https://github.com/yaziciz/GLIMS.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge