Zefan Yang
X-WIN: Building Chest Radiograph World Model via Predictive Sensing
Nov 18, 2025Abstract:Chest X-ray radiography (CXR) is an essential medical imaging technique for disease diagnosis. However, as 2D projectional images, CXRs are limited by structural superposition and hence fail to capture 3D anatomies. This limitation makes representation learning and disease diagnosis challenging. To address this challenge, we propose a novel CXR world model named X-WIN, which distills volumetric knowledge from chest computed tomography (CT) by learning to predict its 2D projections in latent space. The core idea is that a world model with internalized knowledge of 3D anatomical structure can predict CXRs under various transformations in 3D space. During projection prediction, we introduce an affinity-guided contrastive alignment loss that leverages mutual similarities to capture rich, correlated information across projections from the same volume. To improve model adaptability, we incorporate real CXRs into training through masked image modeling and employ a domain classifier to encourage statistically similar representations for real and simulated CXRs. Comprehensive experiments show that X-WIN outperforms existing foundation models on diverse downstream tasks using linear probing and few-shot fine-tuning. X-WIN also demonstrates the ability to render 2D projections for reconstructing a 3D CT volume.
CXR-LT 2024: A MICCAI challenge on long-tailed, multi-label, and zero-shot disease classification from chest X-ray
Jun 09, 2025Abstract:The CXR-LT series is a community-driven initiative designed to enhance lung disease classification using chest X-rays (CXR). It tackles challenges in open long-tailed lung disease classification and enhances the measurability of state-of-the-art techniques. The first event, CXR-LT 2023, aimed to achieve these goals by providing high-quality benchmark CXR data for model development and conducting comprehensive evaluations to identify ongoing issues impacting lung disease classification performance. Building on the success of CXR-LT 2023, the CXR-LT 2024 expands the dataset to 377,110 chest X-rays (CXRs) and 45 disease labels, including 19 new rare disease findings. It also introduces a new focus on zero-shot learning to address limitations identified in the previous event. Specifically, CXR-LT 2024 features three tasks: (i) long-tailed classification on a large, noisy test set, (ii) long-tailed classification on a manually annotated "gold standard" subset, and (iii) zero-shot generalization to five previously unseen disease findings. This paper provides an overview of CXR-LT 2024, detailing the data curation process and consolidating state-of-the-art solutions, including the use of multimodal models for rare disease detection, advanced generative approaches to handle noisy labels, and zero-shot learning strategies for unseen diseases. Additionally, the expanded dataset enhances disease coverage to better represent real-world clinical settings, offering a valuable resource for future research. By synthesizing the insights and innovations of participating teams, we aim to advance the development of clinically realistic and generalizable diagnostic models for chest radiography.
Chest X-ray Foundation Model with Global and Local Representations Integration
Feb 07, 2025Abstract:Chest X-ray (CXR) is the most frequently ordered imaging test, supporting diverse clinical tasks from thoracic disease detection to postoperative monitoring. However, task-specific classification models are limited in scope, require costly labeled data, and lack generalizability to out-of-distribution datasets. To address these challenges, we introduce CheXFound, a self-supervised vision foundation model that learns robust CXR representations and generalizes effectively across a wide range of downstream tasks. We pretrain CheXFound on a curated CXR-1M dataset, comprising over one million unique CXRs from publicly available sources. We propose a Global and Local Representations Integration (GLoRI) module for downstream adaptations, by incorporating disease-specific local features with global image features for enhanced performance in multilabel classification. Our experimental results show that CheXFound outperforms state-of-the-art models in classifying 40 disease findings across different prevalence levels on the CXR-LT 24 dataset and exhibits superior label efficiency on downstream tasks with limited training data. Additionally, CheXFound achieved significant improvements on new tasks with out-of-distribution datasets, including opportunistic cardiovascular disease risk estimation and mortality prediction. These results highlight CheXFound's strong generalization capabilities, enabling diverse adaptations with improved label efficiency. The project source code is publicly available at https://github.com/RPIDIAL/CheXFound.
Cardiovascular Disease Detection from Multi-View Chest X-rays with BI-Mamba
May 28, 2024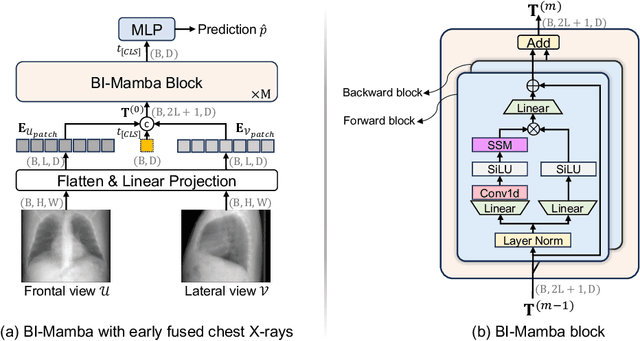
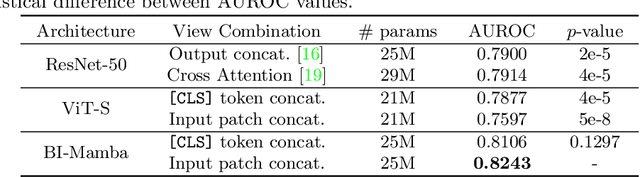

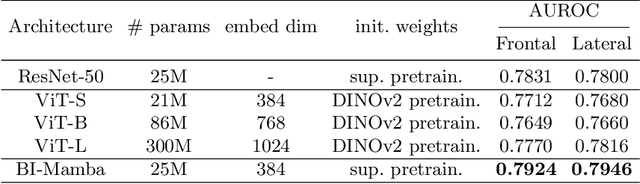
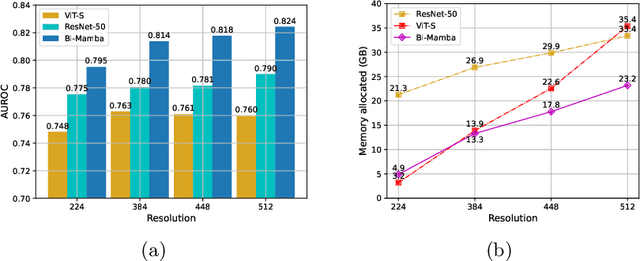
Abstract:Accurate prediction of Cardiovascular disease (CVD) risk in medical imaging is central to effective patient health management. Previous studies have demonstrated that imaging features in computed tomography (CT) can help predict CVD risk. However, CT entails notable radiation exposure, which may result in adverse health effects for patients. In contrast, chest X-ray emits significantly lower levels of radiation, offering a safer option. This rationale motivates our investigation into the feasibility of using chest X-ray for predicting CVD risk. Convolutional Neural Networks (CNNs) and Transformers are two established network architectures for computer-aided diagnosis. However, they struggle to model very high resolution chest X-ray due to the lack of large context modeling power or quadratic time complexity. Inspired by state space sequence models (SSMs), a new class of network architectures with competitive sequence modeling power as Transfomers and linear time complexity, we propose Bidirectional Image Mamba (BI-Mamba) to complement the unidirectional SSMs with opposite directional information. BI-Mamba utilizes parallel forward and backwark blocks to encode longe-range dependencies of multi-view chest X-rays. We conduct extensive experiments on images from 10,395 subjects in National Lung Screening Trail (NLST). Results show that BI-Mamba outperforms ResNet-50 and ViT-S with comparable parameter size, and saves significant amount of GPU memory during training. Besides, BI-Mamba achieves promising performance compared with previous state of the art in CT, unraveling the potential of chest X-ray for CVD risk prediction.
Recurrent Feature Propagation and Edge Skip-Connections for Automatic Abdominal Organ Segmentation
Jan 02, 2022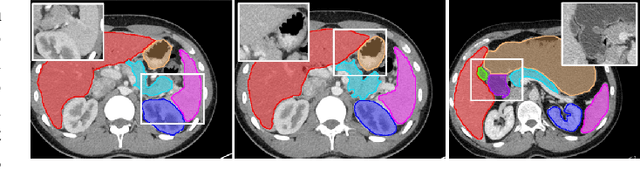
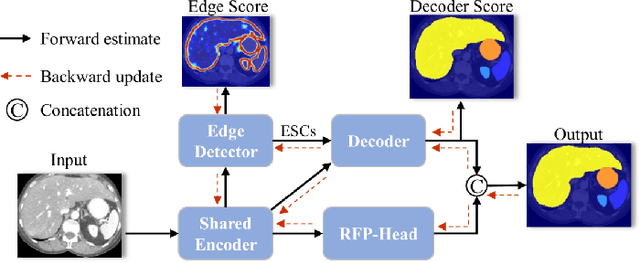
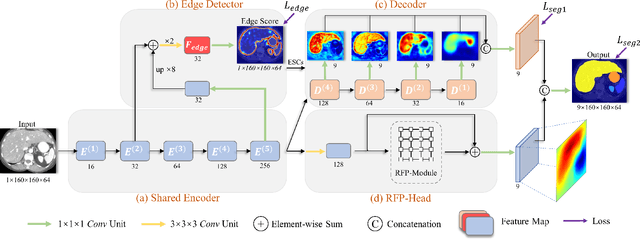
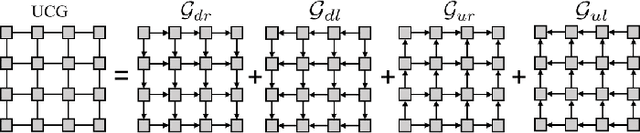
Abstract:Automatic segmentation of abdominal organs in computed tomography (CT) images can support radiation therapy and image-guided surgery workflows. Developing of such automatic solutions remains challenging mainly owing to complex organ interactions and blurry boundaries in CT images. To address these issues, we focus on effective spatial context modeling and explicit edge segmentation priors. Accordingly, we propose a 3D network with four main components trained end-to-end including shared encoder, edge detector, decoder with edge skip-connections (ESCs) and recurrent feature propagation head (RFP-Head). To capture wide-range spatial dependencies, the RFP-Head propagates and harvests local features through directed acyclic graphs (DAGs) formulated with recurrent connections in an efficient slice-wise manner, with regard to spatial arrangement of image units. To leverage edge information, the edge detector learns edge prior knowledge specifically tuned for semantic segmentation by exploiting intermediate features from the encoder with the edge supervision. The ESCs then aggregate the edge knowledge with multi-level decoder features to learn a hierarchy of discriminative features explicitly modeling complementarity between organs' interiors and edges for segmentation. We conduct extensive experiments on two challenging abdominal CT datasets with eight annotated organs. Experimental results show that the proposed network outperforms several state-of-the-art models, especially for the segmentation of small and complicated structures (gallbladder, esophagus, stomach, pancreas and duodenum). The code will be publicly available.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge