Valentin Anklin
HistoCartography: A Toolkit for Graph Analytics in Digital Pathology
Jul 21, 2021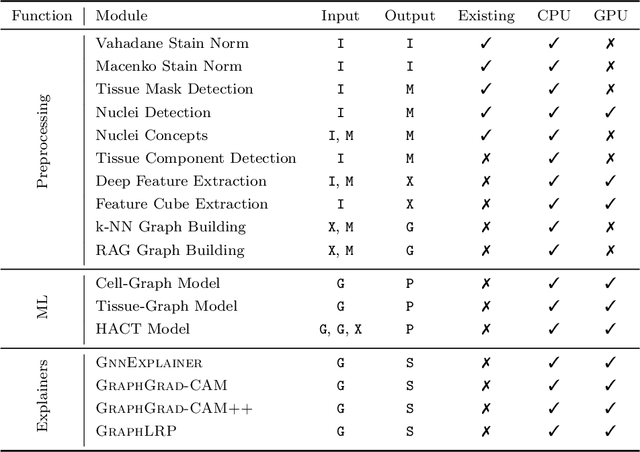
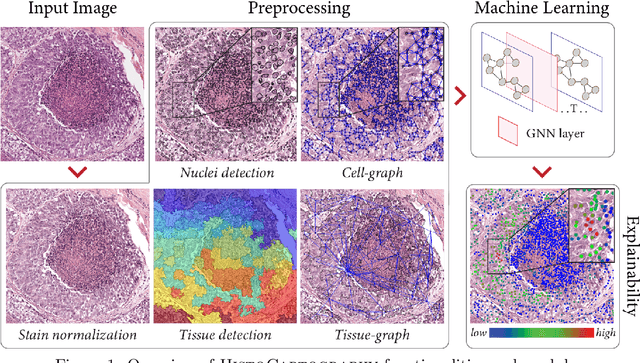
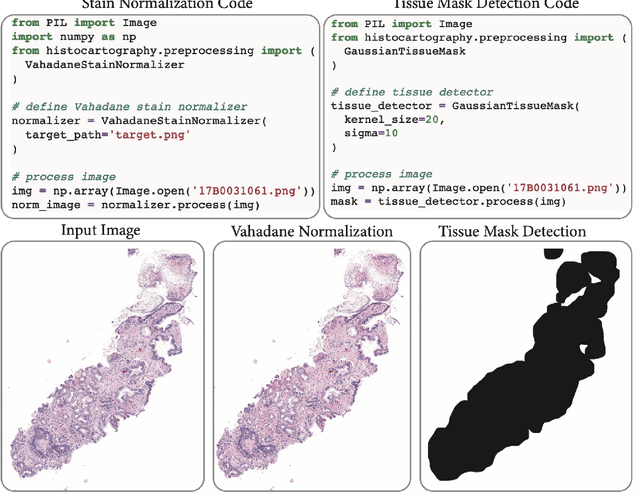
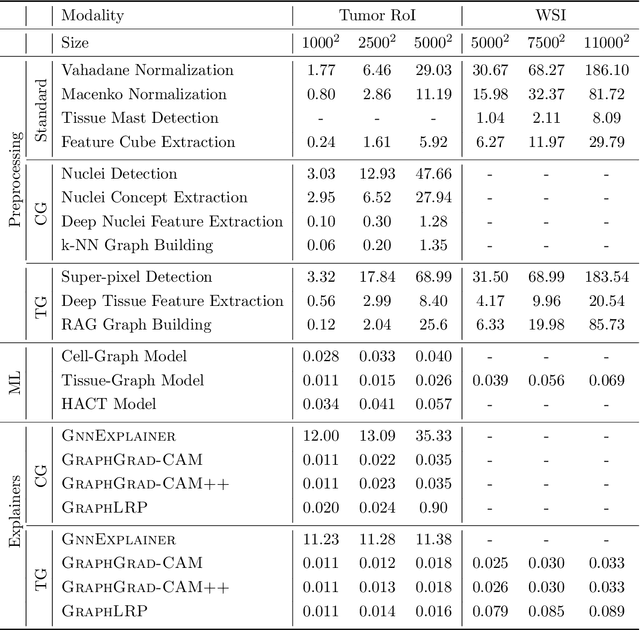
Abstract:Advances in entity-graph based analysis of histopathology images have brought in a new paradigm to describe tissue composition, and learn the tissue structure-to-function relationship. Entity-graphs offer flexible and scalable representations to characterize tissue organization, while allowing the incorporation of prior pathological knowledge to further support model interpretability and explainability. However, entity-graph analysis requires prerequisites for image-to-graph translation and knowledge of state-of-the-art machine learning algorithms applied to graph-structured data, which can potentially hinder their adoption. In this work, we aim to alleviate these issues by developing HistoCartography, a standardized python API with necessary preprocessing, machine learning and explainability tools to facilitate graph-analytics in computational pathology. Further, we have benchmarked the computational time and performance on multiple datasets across different imaging types and histopathology tasks to highlight the applicability of the API for building computational pathology workflows.
Learning Whole-Slide Segmentation from Inexact and Incomplete Labels using Tissue Graphs
Mar 04, 2021Abstract:Segmenting histology images into diagnostically relevant regions is imperative to support timely and reliable decisions by pathologists. To this end, computer-aided techniques have been proposed to delineate relevant regions in scanned histology slides. However, the techniques necessitate task-specific large datasets of annotated pixels, which is tedious, time-consuming, expensive, and infeasible to acquire for many histology tasks. Thus, weakly-supervised semantic segmentation techniques are proposed to utilize weak supervision that is cheaper and quicker to acquire. In this paper, we propose SegGini, a weakly supervised segmentation method using graphs, that can utilize weak multiplex annotations, i.e. inexact and incomplete annotations, to segment arbitrary and large images, scaling from tissue microarray (TMA) to whole slide image (WSI). Formally, SegGini constructs a tissue-graph representation for an input histology image, where the graph nodes depict tissue regions. Then, it performs weakly-supervised segmentation via node classification by using inexact image-level labels, incomplete scribbles, or both. We evaluated SegGini on two public prostate cancer datasets containing TMAs and WSIs. Our method achieved state-of-the-art segmentation performance on both datasets for various annotation settings while being comparable to a pathologist baseline.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge