Theekshana Dissanayake
Patient-independent Epileptic Seizure Prediction using Deep Learning Models
Nov 18, 2020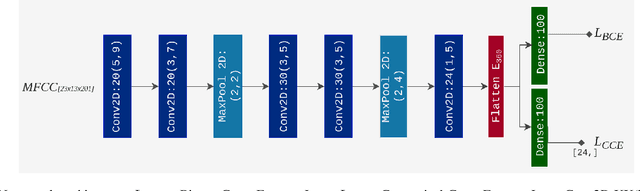
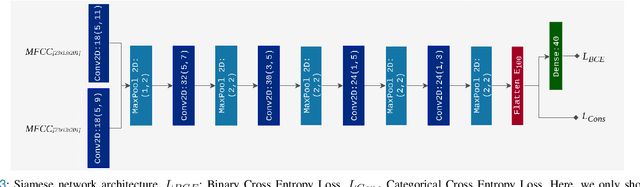
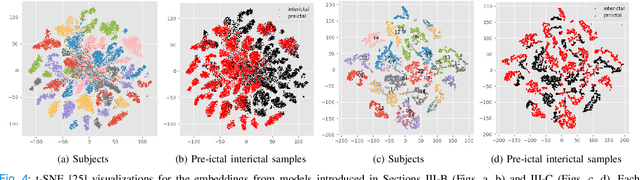
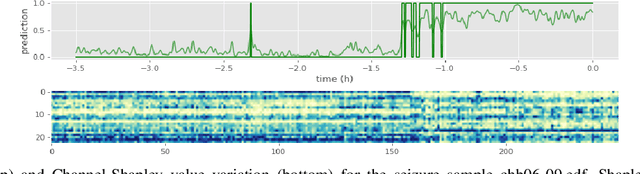
Abstract:Objective: Epilepsy is one of the most prevalent neurological diseases among humans and can lead to severe brain injuries, strokes, and brain tumors. Early detection of seizures can help to mitigate injuries, and can be used to aid the treatment of patients with epilepsy. The purpose of a seizure prediction system is to successfully identify the pre-ictal brain stage, which occurs before a seizure event. Patient-independent seizure prediction models are designed to offer accurate performance across multiple subjects within a dataset, and have been identified as a real-world solution to the seizure prediction problem. However, little attention has been given for designing such models to adapt to the high inter-subject variability in EEG data. Methods: We propose two patient-independent deep learning architectures with different learning strategies that can learn a global function utilizing data from multiple subjects. Results: Proposed models achieve state-of-the-art performance for seizure prediction on the CHB-MIT-EEG dataset, demonstrating 88.81% and 91.54% accuracy respectively. Conclusions: The Siamese model trained on the proposed learning strategy is able to learn patterns related to patient variations in data while predicting seizures. Significance: Our models show superior performance for patient-independent seizure prediction, and the same architecture can be used as a patient-specific classifier after model adaptation. We are the first study that employs model interpretation to understand classifier behavior for the task for seizure prediction, and we also show that the MFCC feature map utilized by our models contains predictive biomarkers related to interictal and pre-ictal brain states.
Domain Generalization in Biosignal Classification
Nov 12, 2020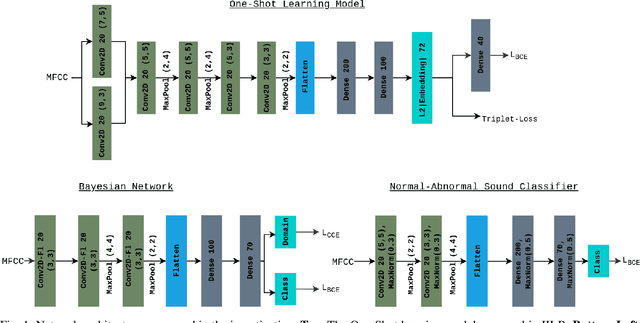
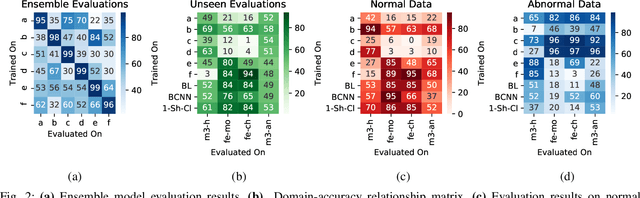
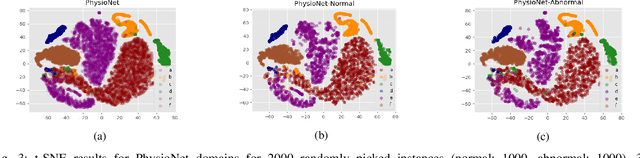
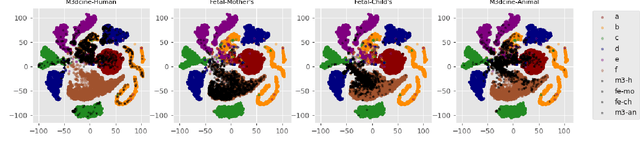
Abstract:Objective: When training machine learning models, we often assume that the training data and evaluation data are sampled from the same distribution. However, this assumption is violated when the model is evaluated on another unseen but similar database, even if that database contains the same classes. This problem is caused by domain-shift and can be solved using two approaches: domain adaptation and domain generalization. Simply, domain adaptation methods can access data from unseen domains during training; whereas in domain generalization, the unseen data is not available during training. Hence, domain generalization concerns models that perform well on inaccessible, domain-shifted data. Method: Our proposed domain generalization method represents an unseen domain using a set of known basis domains, afterwhich we classify the unseen domain using classifier fusion. To demonstrate our system, we employ a collection of heart sound databases that contain normal and abnormal sounds (classes). Results: Our proposed classifier fusion method achieves accuracy gains of up to 16% for four completely unseen domains. Conclusion: Recognizing the complexity induced by the inherent temporal nature of biosignal data, the two-stage method proposed in this study is able to effectively simplify the whole process of domain generalization while demonstrating good results on unseen domains and the adopted basis domains. Significance: To our best knowledge, this is the first study that investigates domain generalization for biosignal data. Our proposed learning strategy can be used to effectively learn domain-relevant features while being aware of the class differences in the data.
Understanding the Importance of Heart Sound Segmentation for Heart Anomaly Detection
May 21, 2020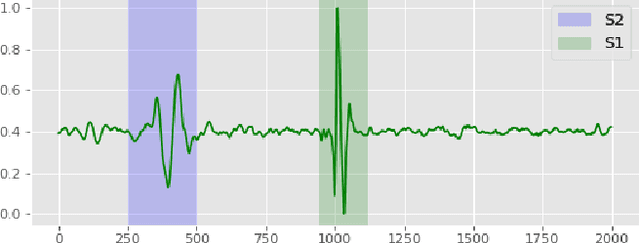
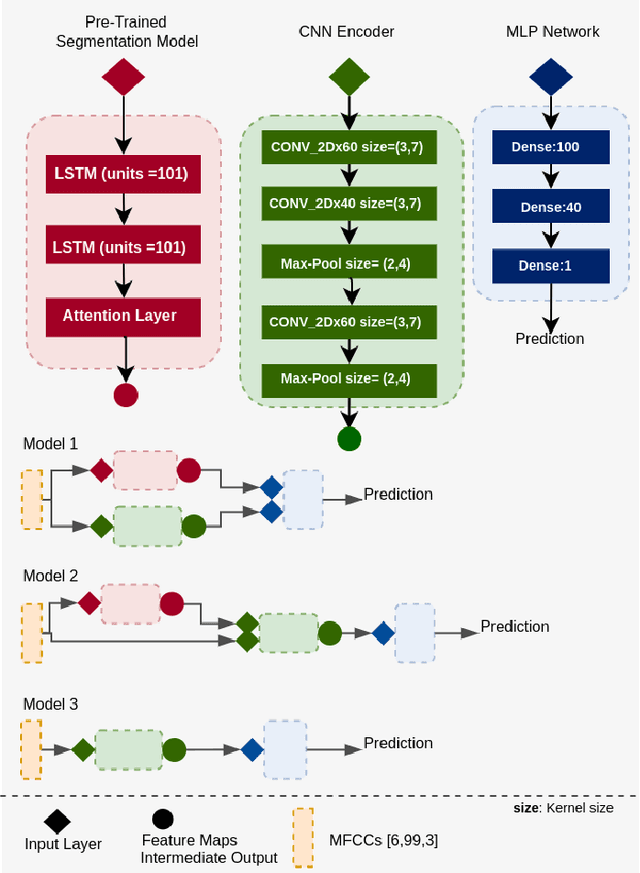


Abstract:Traditionally, abnormal heart sound classification is framed as a three-stage process. The first stage involves segmenting the phonocardiogram to detect fundamental heart sounds; after which features are extracted and classification is performed. Some researchers in the field argue the segmentation step is an unwanted computational burden, whereas others embrace it as a prior step to feature extraction. When comparing accuracies achieved by studies that have segmented heart sounds before analysis with those who have overlooked that step, the question of whether to segment heart sounds before feature extraction is still open. In this study, we explicitly examine the importance of heart sound segmentation as a prior step for heart sound classification, and then seek to apply the obtained insights to propose a robust classifier for abnormal heart sound detection. Furthermore, recognizing the pressing need for explainable Artificial Intelligence (AI) models in the medical domain, we also unveil hidden representations learned by the classifier using model interpretation techniques. Experimental results demonstrate that the segmentation plays an essential role in abnormal heart sound classification. Our new classifier is also shown to be robust, stable and most importantly, explainable, with an accuracy of almost 100% on the widely used PhysioNet dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge