Tej Phatak
Attention-based Multi-scale Gated Recurrent Encoder with Novel Correlation Loss for COVID-19 Progression Prediction
Jul 18, 2021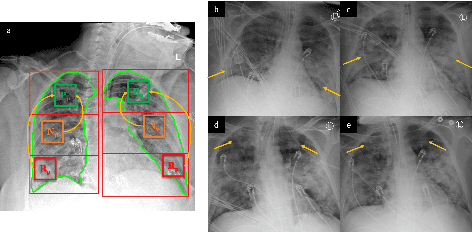


Abstract:COVID-19 image analysis has mostly focused on diagnostic tasks using single timepoint scans acquired upon disease presentation or admission. We present a deep learning-based approach to predict lung infiltrate progression from serial chest radiographs (CXRs) of COVID-19 patients. Our method first utilizes convolutional neural networks (CNNs) for feature extraction from patches within the concerned lung zone, and also from neighboring and remote boundary regions. The framework further incorporates a multi-scale Gated Recurrent Unit (GRU) with a correlation module for effective predictions. The GRU accepts CNN feature vectors from three different areas as input and generates a fused representation. The correlation module attempts to minimize the correlation loss between hidden representations of concerned and neighboring area feature vectors, while maximizing the loss between the same from concerned and remote regions. Further, we employ an attention module over the output hidden states of each encoder timepoint to generate a context vector. This vector is used as an input to a decoder module to predict patch severity grades at a future timepoint. Finally, we ensemble the patch classification scores to calculate patient-wise grades. Specifically, our framework predicts zone-wise disease severity for a patient on a given day by learning representations from the previous temporal CXRs. Our novel multi-institutional dataset comprises sequential CXR scans from N=93 patients. Our approach outperforms transfer learning and radiomic feature-based baseline approaches on this dataset.
Attention based CNN-LSTM Network for Pulmonary Embolism Prediction on Chest Computed Tomography Pulmonary Angiograms
Jul 13, 2021



Abstract:With more than 60,000 deaths annually in the United States, Pulmonary Embolism (PE) is among the most fatal cardiovascular diseases. It is caused by an artery blockage in the lung; confirming its presence is time-consuming and is prone to over-diagnosis. The utilization of automated PE detection systems is critical for diagnostic accuracy and efficiency. In this study we propose a two-stage attention-based CNN-LSTM network for predicting PE, its associated type (chronic, acute) and corresponding location (leftsided, rightsided or central) on computed tomography (CT) examinations. We trained our model on the largest available public Computed Tomography Pulmonary Angiogram PE dataset (RSNA-STR Pulmonary Embolism CT (RSPECT) Dataset, N=7279 CT studies) and tested it on an in-house curated dataset of N=106 studies. Our framework mirrors the radiologic diagnostic process via a multi-slice approach so that the accuracy and pathologic sequela of true pulmonary emboli may be meticulously assessed, enabling physicians to better appraise the morbidity of a PE when present. Our proposed method outperformed a baseline CNN classifier and a single-stage CNN-LSTM network, achieving an AUC of 0.95 on the test set for detecting the presence of PE in the study.
Predicting Mechanical Ventilation Requirement and Mortality in COVID-19 using Radiomics and Deep Learning on Chest Radiographs: A Multi-Institutional Study
Jul 15, 2020
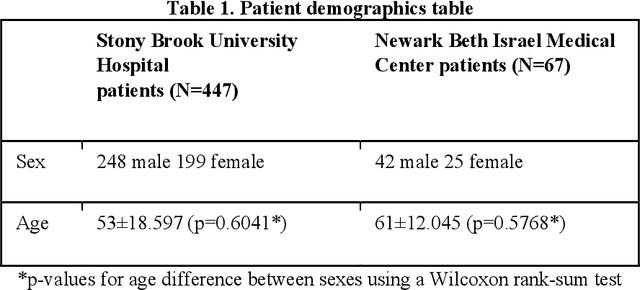


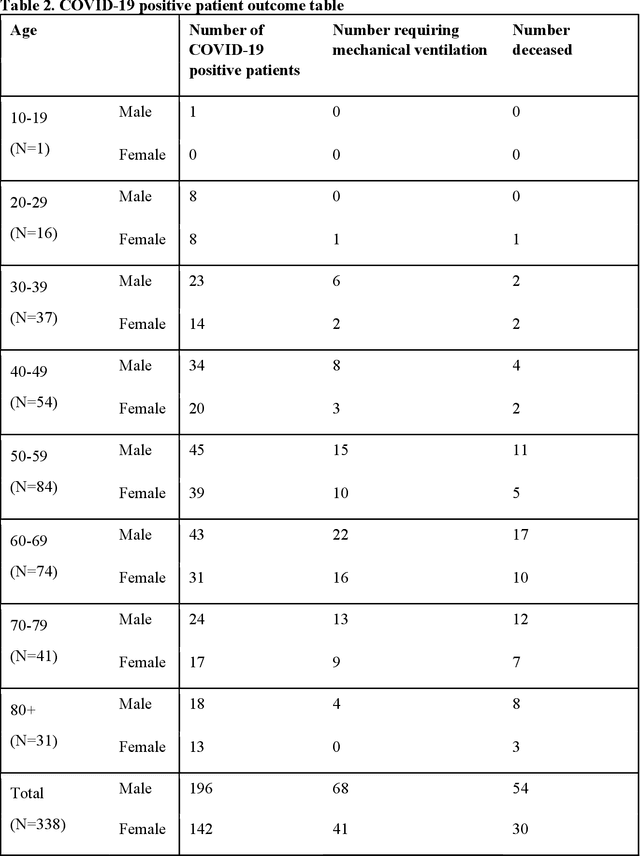
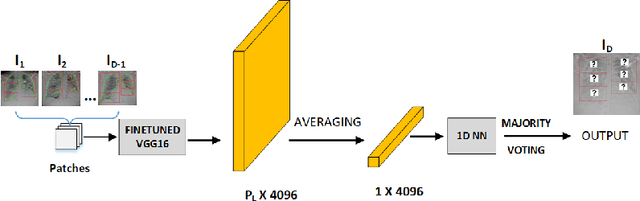
Abstract:Objectives: To predict mechanical ventilation requirement and mortality using computational modeling of chest radiographs (CXR) for coronavirus disease 2019 (COVID-19) patients. We also investigate the relative advantages of deep learning (DL), radiomics, and DL of radiomic-embedded feature maps in predicting these outcomes. Methods: This two-center, retrospective study analyzed deidentified CXRs taken from 514 patients suspected of COVID-19 infection on presentation at Stony Brook University Hospital (SBUH) and Newark Beth Israel Medical Center (NBIMC) between the months of March and June 2020. A DL segmentation pipeline was developed to generate masks for both lung fields and artifacts for each CXR. Machine learning classifiers to predict mechanical ventilation requirement and mortality were trained and evaluated on 353 baseline CXRs taken from COVID-19 positive patients. A novel radiomic embedding framework is also explored for outcome prediction. Results: Classification models for mechanical ventilation requirement (test N=154) and mortality (test N=190) had AUCs of up to 0.904 and 0.936, respectively. We also found that the inclusion of radiomic-embedded maps improved DL model predictions of clinical outcomes. Conclusions: We demonstrate the potential for computerized analysis of baseline CXR in predicting disease outcomes in COVID-19 patients. Our results also suggest that radiomic embedding improves DL models in medical image analysis, a technique that might be explored further in other pathologies. The models proposed in this study and the prognostic information they provide, complementary to other clinical data, might be used to aid physician decision making and resource allocation during the COVID-19 pandemic.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge