Tammie L. S. Benzinger
Analysing heterogeneity in Alzheimer Disease using multimodal normative modelling on ATN biomarkers
Apr 04, 2024Abstract:Alzheimer Disease (AD) is a multi-faceted disorder, with each modality providing unique and complementary info about AD. In this study, we used a deep-learning based multimodal normative model to assess the heterogeneity in regional brain patterns for ATN (amyloid-tau-neurodegeneration) biomarkers. We selected discovery (n = 665) and replication (n = 430) cohorts with simultaneous availability of ATN biomarkers: Florbetapir amyloid, Flortaucipir tau and T1-weighted MRI (magnetic resonance imaging) imaging. A multimodal variational autoencoder (conditioned on age and sex) was used as a normative model to learn the multimodal regional brain patterns of a cognitively unimpaired (CU) control group. The trained model was applied on individuals on the ADS (AD Spectrum) to estimate their deviations (Z-scores) from the normative distribution, resulting in a Z-score regional deviation map per ADS individual per modality. ADS individuals with moderate or severe dementia showed higher proportion of regional outliers for each modality as well as more dissimilarity in modality-specific regional outlier patterns compared to ADS individuals with early or mild dementia. DSI was associated with the progressive stages of dementia, (ii) showed significant associations with neuropsychological composite scores and (iii) related to the longitudinal risk of CDR progression. Findings were reproducible in both discovery and replication cohorts. Our is the first study to examine the heterogeneity in AD through the lens of multiple neuroimaging modalities (ATN), based on distinct or overlapping patterns of regional outlier deviations. Regional MRI and tau outliers were more heterogenous than regional amyloid outliers. DSI has the potential to be an individual patient metric of neurodegeneration that can help in clinical decision making and monitoring patient response for anti-amyloid treatments.
Gene-SGAN: a method for discovering disease subtypes with imaging and genetic signatures via multi-view weakly-supervised deep clustering
Jan 25, 2023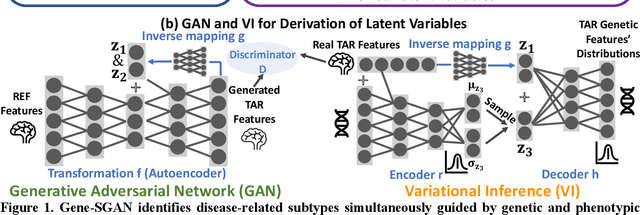
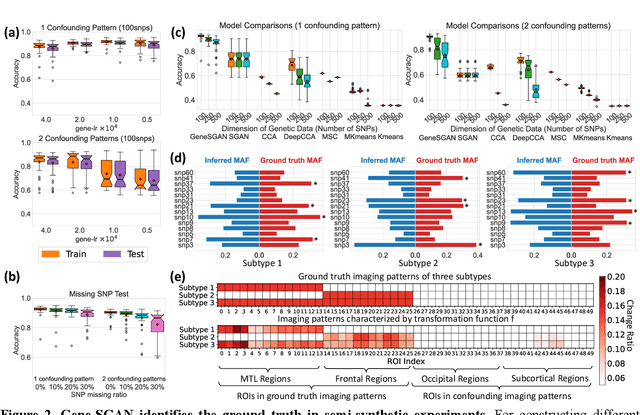
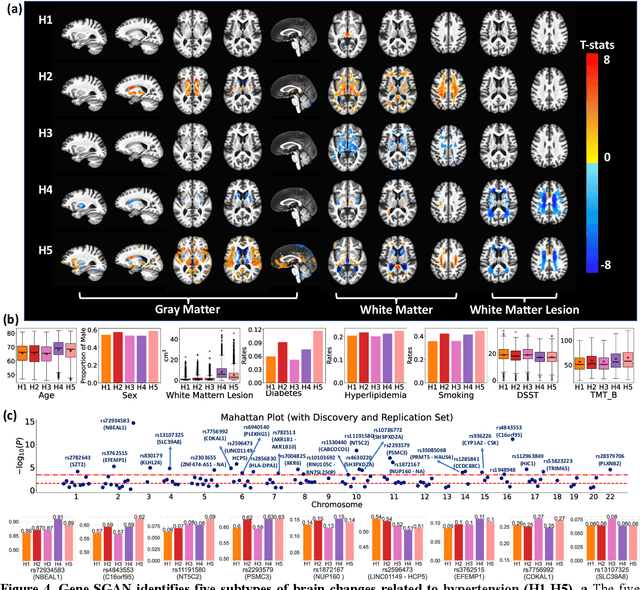
Abstract:Disease heterogeneity has been a critical challenge for precision diagnosis and treatment, especially in neurologic and neuropsychiatric diseases. Many diseases can display multiple distinct brain phenotypes across individuals, potentially reflecting disease subtypes that can be captured using MRI and machine learning methods. However, biological interpretability and treatment relevance are limited if the derived subtypes are not associated with genetic drivers or susceptibility factors. Herein, we describe Gene-SGAN - a multi-view, weakly-supervised deep clustering method - which dissects disease heterogeneity by jointly considering phenotypic and genetic data, thereby conferring genetic correlations to the disease subtypes and associated endophenotypic signatures. We first validate the generalizability, interpretability, and robustness of Gene-SGAN in semi-synthetic experiments. We then demonstrate its application to real multi-site datasets from 28,858 individuals, deriving subtypes of Alzheimer's disease and brain endophenotypes associated with hypertension, from MRI and SNP data. Derived brain phenotypes displayed significant differences in neuroanatomical patterns, genetic determinants, biological and clinical biomarkers, indicating potentially distinct underlying neuropathologic processes, genetic drivers, and susceptibility factors. Overall, Gene-SGAN is broadly applicable to disease subtyping and endophenotype discovery, and is herein tested on disease-related, genetically-driven neuroimaging phenotypes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge