Sven Koehler
Self-supervised motion descriptor for cardiac phase detection in 4D CMR based on discrete vector field estimations
Sep 18, 2022
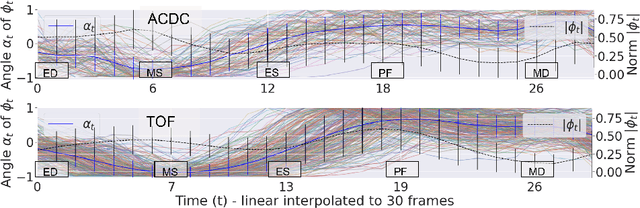
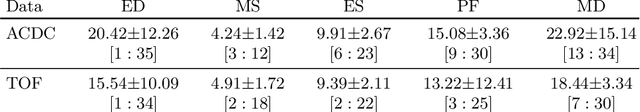
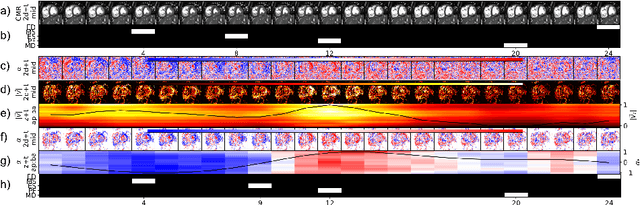
Abstract:Cardiac magnetic resonance (CMR) sequences visualise the cardiac function voxel-wise over time. Simultaneously, deep learning-based deformable image registration is able to estimate discrete vector fields which warp one time step of a CMR sequence to the following in a self-supervised manner. However, despite the rich source of information included in these 3D+t vector fields, a standardised interpretation is challenging and the clinical applications remain limited so far. In this work, we show how to efficiently use a deformable vector field to describe the underlying dynamic process of a cardiac cycle in form of a derived 1D motion descriptor. Additionally, based on the expected cardiovascular physiological properties of a contracting or relaxing ventricle, we define a set of rules that enables the identification of five cardiovascular phases including the end-systole (ES) and end-diastole (ED) without the usage of labels. We evaluate the plausibility of the motion descriptor on two challenging multi-disease, -center, -scanner short-axis CMR datasets. First, by reporting quantitative measures such as the periodic frame difference for the extracted phases. Second, by comparing qualitatively the general pattern when we temporally resample and align the motion descriptors of all instances across both datasets. The average periodic frame difference for the ED, ES key phases of our approach is $0.80\pm{0.85}$, $0.69\pm{0.79}$ which is slightly better than the inter-observer variability ($1.07\pm{0.86}$, $0.91\pm{1.6}$) and the supervised baseline method ($1.18\pm{1.91}$, $1.21\pm{1.78}$). Code and labels will be made available on our GitHub repository. https://github.com/Cardio-AI/cmr-phase-detection
Comparison of Evaluation Metrics for Landmark Detection in CMR Images
Jan 28, 2022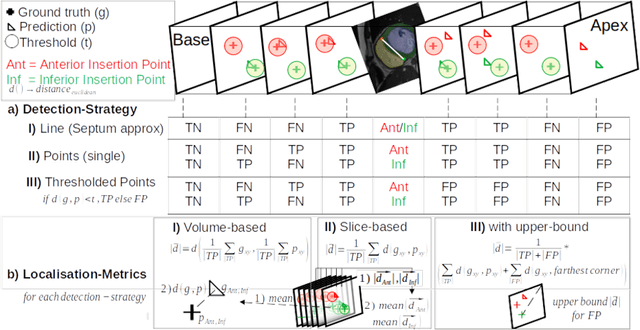

Abstract:Cardiac Magnetic Resonance (CMR) images are widely used for cardiac diagnosis and ventricular assessment. Extracting specific landmarks like the right ventricular insertion points is of importance for spatial alignment and 3D modeling. The automatic detection of such landmarks has been tackled by multiple groups using Deep Learning, but relatively little attention has been paid to the failure cases of evaluation metrics in this field. In this work, we extended the public ACDC dataset with additional labels of the right ventricular insertion points and compare different variants of a heatmap-based landmark detection pipeline. In this comparison, we demonstrate very likely pitfalls of apparently simple detection and localisation metrics which highlights the importance of a clear detection strategy and the definition of an upper limit for localisation-based metrics. Our preliminary results indicate that a combination of different metrics is necessary, as they yield different winners for method comparison. Additionally, they highlight the need of a comprehensive metric description and evaluation standardisation, especially for the error cases where no metrics could be computed or where no lower/upper boundary of a metric exists. Code and labels: https://github.com/Cardio-AI/rvip_landmark_detection
Mutually improved endoscopic image synthesis and landmark detection in unpaired image-to-image translation
Jul 14, 2021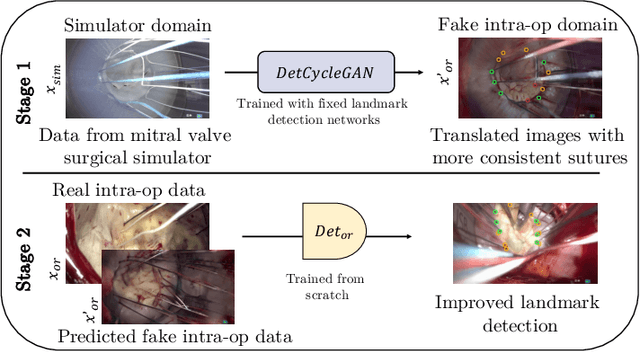
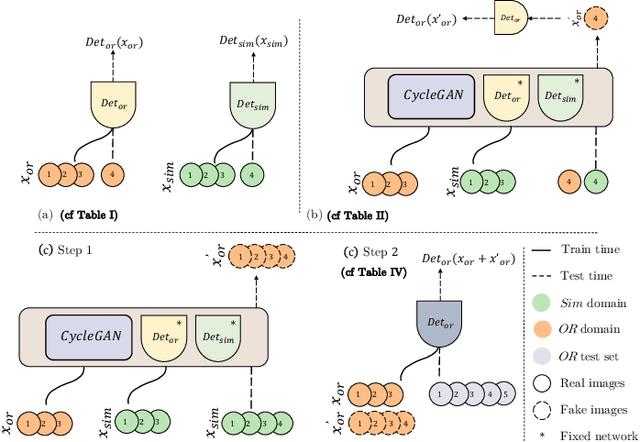
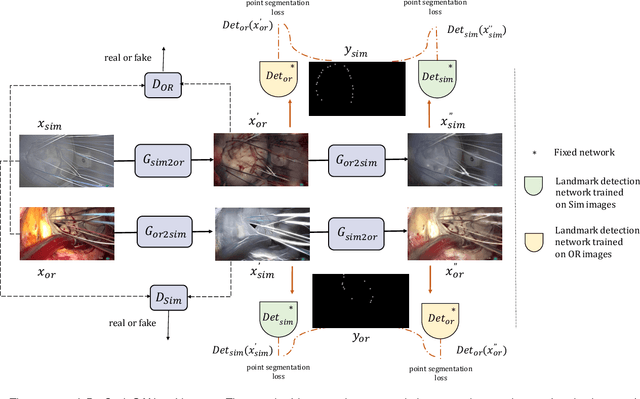
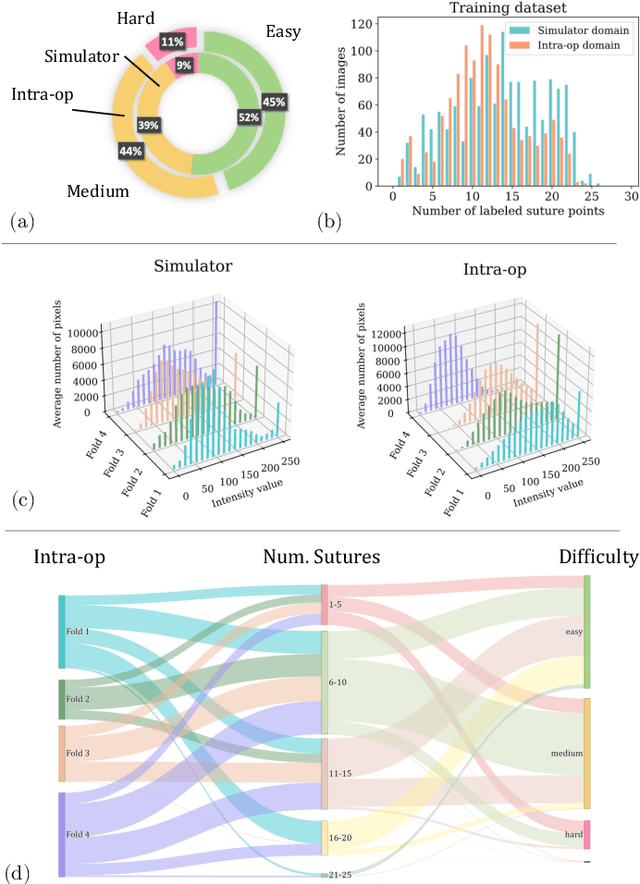
Abstract:The CycleGAN framework allows for unsupervised image-to-image translation of unpaired data. In a scenario of surgical training on a physical surgical simulator, this method can be used to transform endoscopic images of phantoms into images which more closely resemble the intra-operative appearance of the same surgical target structure. This can be viewed as a novel augmented reality approach, which we coined Hyperrealism in previous work. In this use case, it is of paramount importance to display objects like needles, sutures or instruments consistent in both domains while altering the style to a more tissue-like appearance. Segmentation of these objects would allow for a direct transfer, however, contouring of these, partly tiny and thin foreground objects is cumbersome and perhaps inaccurate. Instead, we propose to use landmark detection on the points when sutures pass into the tissue. This objective is directly incorporated into a CycleGAN framework by treating the performance of pre-trained detector models as an additional optimization goal. We show that a task defined on these sparse landmark labels improves consistency of synthesis by the generator network in both domains. Comparing a baseline CycleGAN architecture to our proposed extension (DetCycleGAN), mean precision (PPV) improved by +61.32, mean sensitivity (TPR) by +37.91, and mean F1 score by +0.4743. Furthermore, it could be shown that by dataset fusion, generated intra-operative images can be leveraged as additional training data for the detection network itself. The data is released within the scope of the AdaptOR MICCAI Challenge 2021 at https://adaptor2021.github.io/, and code at https://github.com/Cardio-AI/detcyclegan_pytorch.
Unsupervised Domain Adaptation from Axial to Short-Axis Multi-Slice Cardiac MR Images by Incorporating Pretrained Task Networks
Jan 20, 2021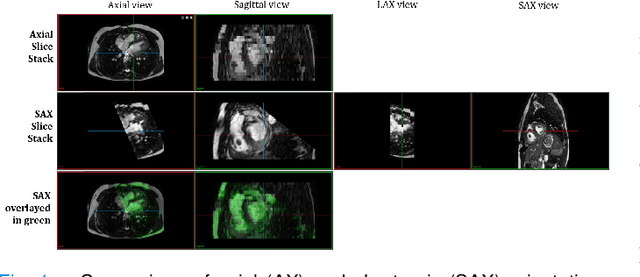
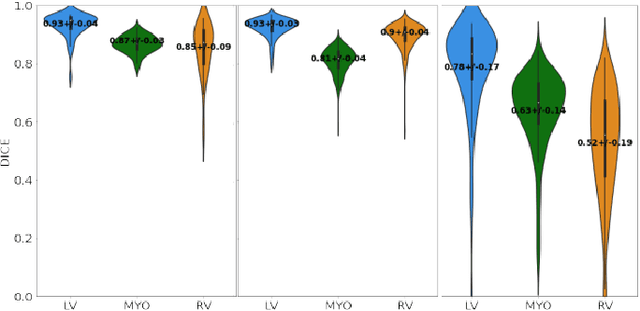
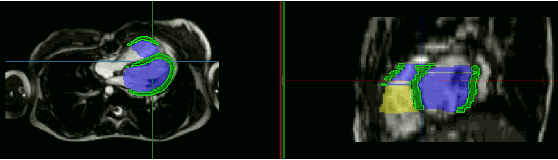
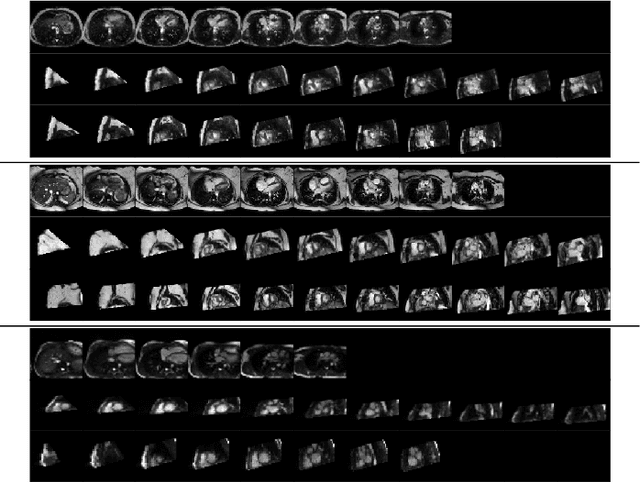
Abstract:Anisotropic multi-slice Cardiac Magnetic Resonance (CMR) Images are conventionally acquired in patient-specific short-axis (SAX) orientation. In specific cardiovascular diseases that affect right ventricular (RV) morphology, acquisitions in standard axial (AX) orientation are preferred by some investigators, due to potential superiority in RV volume measurement for treatment planning. Unfortunately, due to the rare occurrence of these diseases, data in this domain is scarce. Recent research in deep learning-based methods mainly focused on SAX CMR images and they had proven to be very successful. In this work, we show that there is a considerable domain shift between AX and SAX images, and therefore, direct application of existing models yield sub-optimal results on AX samples. We propose a novel unsupervised domain adaptation approach, which uses task-related probabilities in an attention mechanism. Beyond that, cycle consistency is imposed on the learned patient-individual 3D rigid transformation to improve stability when automatically re-sampling the AX images to SAX orientations. The network was trained on 122 registered 3D AX-SAX CMR volume pairs from a multi-centric patient cohort. A mean 3D Dice of $0.86\pm{0.06}$ for the left ventricle, $0.65\pm{0.08}$ for the myocardium, and $0.77\pm{0.10}$ for the right ventricle could be achieved. This is an improvement of $25\%$ in Dice for RV in comparison to direct application on axial slices. To conclude, our pre-trained task module has neither seen CMR images nor labels from the target domain, but is able to segment them after the domain gap is reduced. Code: https://github.com/Cardio-AI/3d-mri-domain-adaptation
Heatmap-based 2D Landmark Detection with a Varying Number of Landmarks
Jan 07, 2021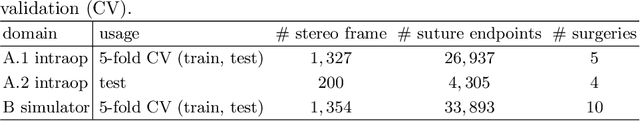
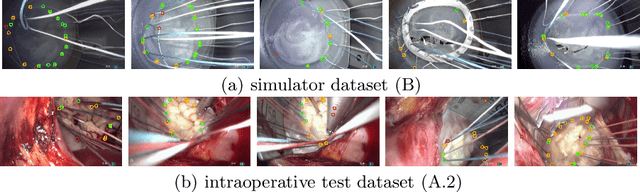
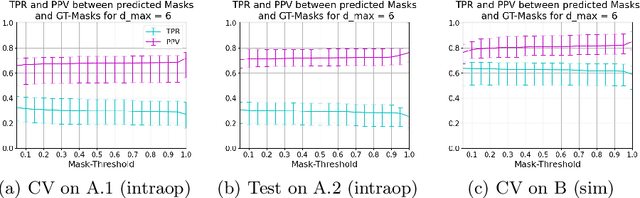
Abstract:Mitral valve repair is a surgery to restore the function of the mitral valve. To achieve this, a prosthetic ring is sewed onto the mitral annulus. Analyzing the sutures, which are punctured through the annulus for ring implantation, can be useful in surgical skill assessment, for quantitative surgery and for positioning a virtual prosthetic ring model in the scene via augmented reality. This work presents a neural network approach which detects the sutures in endoscopic images of mitral valve repair and therefore solves a landmark detection problem with varying amount of landmarks, as opposed to most other existing deep learning-based landmark detection approaches. The neural network is trained separately on two data collections from different domains with the same architecture and hyperparameter settings. The datasets consist of more than 1,300 stereo frame pairs each, with a total over 60,000 annotated landmarks. The proposed heatmap-based neural network achieves a mean positive predictive value (PPV) of 66.68$\pm$4.67% and a mean true positive rate (TPR) of 24.45$\pm$5.06% on the intraoperative test dataset and a mean PPV of 81.50\pm5.77\% and a mean TPR of 61.60$\pm$6.11% on a dataset recorded during surgical simulation. The best detection results are achieved when the camera is positioned above the mitral valve with good illumination. A detection from a sideward view is also possible if the mitral valve is well perceptible.
How well do U-Net-based segmentation trained on adult cardiac magnetic resonance imaging data generalise to rare congenital heart diseases for surgical planning?
Feb 10, 2020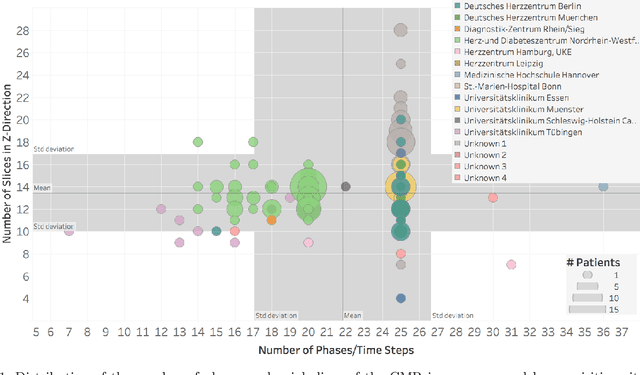
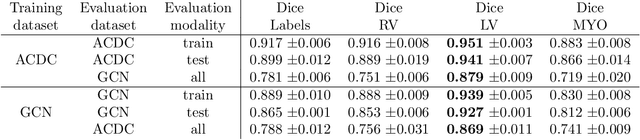


Abstract:Planning the optimal time of intervention for pulmonary valve replacement surgery in patients with the congenital heart disease Tetralogy of Fallot (TOF) is mainly based on ventricular volume and function according to current guidelines. Both of these two biomarkers are most reliably assessed by segmentation of 3D cardiac magnetic resonance (CMR) images. In several grand challenges in the last years, U-Net architectures have shown impressive results on the provided data. However, in clinical practice, data sets are more diverse considering individual pathologies and image properties derived from different scanner properties. Additionally, specific training data for complex rare diseases like TOF is scarce. For this work, 1) we assessed the accuracy gap when using a publicly available labelled data set (the Automatic Cardiac Diagnosis Challenge (ACDC) data set) for training and subsequent applying it to CMR data of TOF patients and vice versa and 2) whether we can achieve similar results when applying the model to a more heterogeneous data base. Multiple deep learning models were trained with four-fold cross validation. Afterwards they were evaluated on the respective unseen CMR images from the other collection. Our results confirm that current deep learning models can achieve excellent results (left ventricle dice of $0.951\pm{0.003}$/$0.941\pm{0.007}$ train/validation) within a single data collection. But once they are applied to other pathologies, it becomes apparent how much they overfit to the training pathologies (dice score drops between $0.072\pm{0.001}$ for the left and $0.165\pm{0.001}$ for the right ventricle).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge