Stephan Lau
Australian Institute for Machine Learning, South Australian Health and Medical Research Institute
Segmentation method for cerebral blood vessels from MRA using hysteresis
Mar 09, 2023Abstract:Segmentation of cerebral blood vessels from Magnetic Resonance Imaging (MRI) is an open problem that could be solved with deep learning (DL). However, annotated data for training is often scarce. Due to the absence of open-source tools, we aim to develop a classical segmentation method that generates vessel ground truth from Magnetic Resonance Angiography for DL training of segmentation across a variety of modalities. The method combines size-specific Hessian filters, hysteresis thresholding and connected component correction. The optimal choice of processing steps was evaluated with a blinded scoring by a clinician using 24 3D images. The results show that all method steps are necessary to produce the highest (14.2/15) vessel segmentation quality score. Omitting the connected component correction caused the largest quality loss. The method, which is available on GitHub, can be used to train DL models for vessel segmentation.
Mutual information neural estimation for unsupervised multi-modal registration of brain images
Jan 25, 2022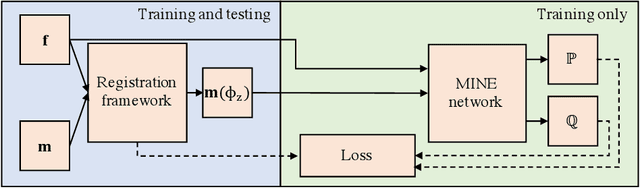
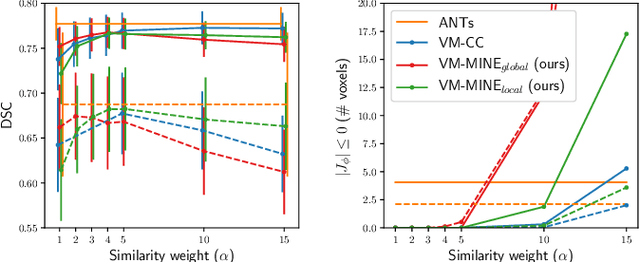
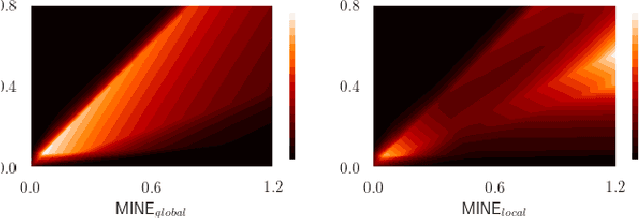

Abstract:Many applications in image-guided surgery and therapy require fast and reliable non-linear, multi-modal image registration. Recently proposed unsupervised deep learning-based registration methods have demonstrated superior performance compared to iterative methods in just a fraction of the time. Most of the learning-based methods have focused on mono-modal image registration. The extension to multi-modal registration depends on the use of an appropriate similarity function, such as the mutual information (MI). We propose guiding the training of a deep learning-based registration method with MI estimation between an image-pair in an end-to-end trainable network. Our results show that a small, 2-layer network produces competitive results in both mono- and multimodal registration, with sub-second run-times. Comparisons to both iterative and deep learning-based methods show that our MI-based method produces topologically and qualitatively superior results with an extremely low rate of non-diffeomorphic transformations. Real-time clinical application will benefit from a better visual matching of anatomical structures and less registration failures/outliers.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge