Shota Nakamura
Micro CT Image-Assisted Cross Modality Super-Resolution of Clinical CT Images Utilizing Synthesized Training Dataset
Oct 20, 2020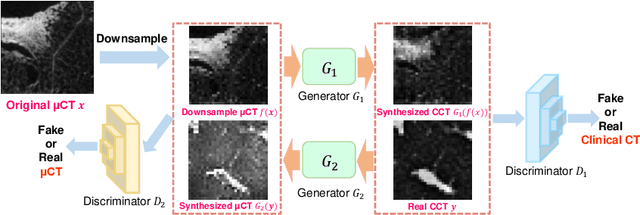
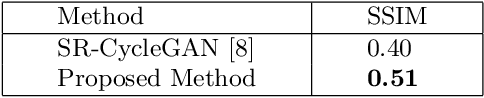
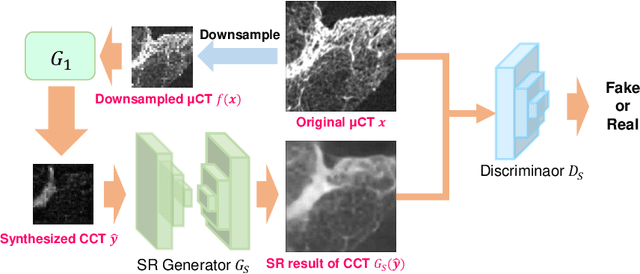

Abstract:This paper proposes a novel, unsupervised super-resolution (SR) approach for performing the SR of a clinical CT into the resolution level of a micro CT ($\mu$CT). The precise non-invasive diagnosis of lung cancer typically utilizes clinical CT data. Due to the resolution limitations of clinical CT (about $0.5 \times 0.5 \times 0.5$ mm$^3$), it is difficult to obtain enough pathological information such as the invasion area at alveoli level. On the other hand, $\mu$CT scanning allows the acquisition of volumes of lung specimens with much higher resolution ($50 \times 50 \times 50 \mu {\rm m}^3$ or higher). Thus, super-resolution of clinical CT volume may be helpful for diagnosis of lung cancer. Typical SR methods require aligned pairs of low-resolution (LR) and high-resolution (HR) images for training. Unfortunately, obtaining paired clinical CT and $\mu$CT volumes of human lung tissues is infeasible. Unsupervised SR methods are required that do not need paired LR and HR images. In this paper, we create corresponding clinical CT-$\mu$CT pairs by simulating clinical CT images from $\mu$CT images by modified CycleGAN. After this, we use simulated clinical CT-$\mu$CT image pairs to train an SR network based on SRGAN. Finally, we use the trained SR network to perform SR of the clinical CT images. We compare our proposed method with another unsupervised SR method for clinical CT images named SR-CycleGAN. Experimental results demonstrate that the proposed method can successfully perform SR of clinical CT images of lung cancer patients with $\mu$CT level resolution, and quantitatively and qualitatively outperformed conventional method (SR-CycleGAN), improving the SSIM (structure similarity) form 0.40 to 0.51.
Multi-modality super-resolution loss for GAN-based super-resolution of clinical CT images using micro CT image database
Dec 30, 2019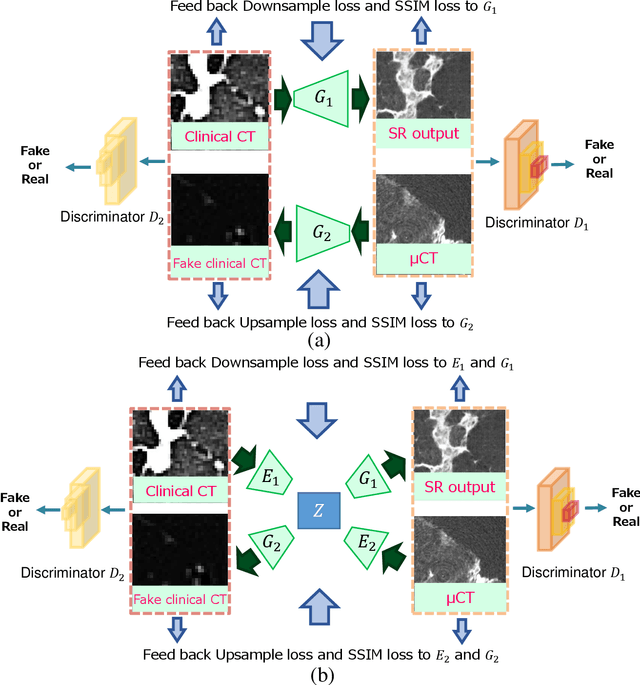
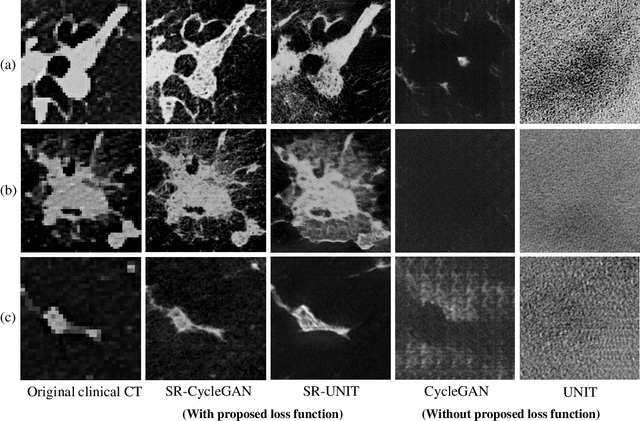
Abstract:This paper newly introduces multi-modality loss function for GAN-based super-resolution that can maintain image structure and intensity on unpaired training dataset of clinical CT and micro CT volumes. Precise non-invasive diagnosis of lung cancer mainly utilizes 3D multidetector computed-tomography (CT) data. On the other hand, we can take micro CT images of resected lung specimen in 50 micro meter or higher resolution. However, micro CT scanning cannot be applied to living human imaging. For obtaining highly detailed information such as cancer invasion area from pre-operative clinical CT volumes of lung cancer patients, super-resolution (SR) of clinical CT volumes to $\mu$CT level might be one of substitutive solutions. While most SR methods require paired low- and high-resolution images for training, it is infeasible to obtain precisely paired clinical CT and micro CT volumes. We aim to propose unpaired SR approaches for clincial CT using micro CT images based on unpaired image translation methods such as CycleGAN or UNIT. Since clinical CT and micro CT are very different in structure and intensity, direct application of GAN-based unpaired image translation methods in super-resolution tends to generate arbitrary images. Aiming to solve this problem, we propose new loss function called multi-modality loss function to maintain the similarity of input images and corresponding output images in super-resolution task. Experimental results demonstrated that the newly proposed loss function made CycleGAN and UNIT to successfully perform SR of clinical CT images of lung cancer patients into micro CT level resolution, while original CycleGAN and UNIT failed in super-resolution.
Unsupervised Segmentation of 3D Medical Images Based on Clustering and Deep Representation Learning
Apr 11, 2018Abstract:This paper presents a novel unsupervised segmentation method for 3D medical images. Convolutional neural networks (CNNs) have brought significant advances in image segmentation. However, most of the recent methods rely on supervised learning, which requires large amounts of manually annotated data. Thus, it is challenging for these methods to cope with the growing amount of medical images. This paper proposes a unified approach to unsupervised deep representation learning and clustering for segmentation. Our proposed method consists of two phases. In the first phase, we learn deep feature representations of training patches from a target image using joint unsupervised learning (JULE) that alternately clusters representations generated by a CNN and updates the CNN parameters using cluster labels as supervisory signals. We extend JULE to 3D medical images by utilizing 3D convolutions throughout the CNN architecture. In the second phase, we apply k-means to the deep representations from the trained CNN and then project cluster labels to the target image in order to obtain the fully segmented image. We evaluated our methods on three images of lung cancer specimens scanned with micro-computed tomography (micro-CT). The automatic segmentation of pathological regions in micro-CT could further contribute to the pathological examination process. Hence, we aim to automatically divide each image into the regions of invasive carcinoma, noninvasive carcinoma, and normal tissue. Our experiments show the potential abilities of unsupervised deep representation learning for medical image segmentation.
* This paper was presented at SPIE Medical Imaging 2018, Houston, TX, USA
Unsupervised Pathology Image Segmentation Using Representation Learning with Spherical K-means
Apr 11, 2018Abstract:This paper presents a novel method for unsupervised segmentation of pathology images. Staging of lung cancer is a major factor of prognosis. Measuring the maximum dimensions of the invasive component in a pathology images is an essential task. Therefore, image segmentation methods for visualizing the extent of invasive and noninvasive components on pathology images could support pathological examination. However, it is challenging for most of the recent segmentation methods that rely on supervised learning to cope with unlabeled pathology images. In this paper, we propose a unified approach to unsupervised representation learning and clustering for pathology image segmentation. Our method consists of two phases. In the first phase, we learn feature representations of training patches from a target image using the spherical k-means. The purpose of this phase is to obtain cluster centroids which could be used as filters for feature extraction. In the second phase, we apply conventional k-means to the representations extracted by the centroids and then project cluster labels to the target images. We evaluated our methods on pathology images of lung cancer specimen. Our experiments showed that the proposed method outperforms traditional k-means segmentation and the multithreshold Otsu method both quantitatively and qualitatively with an improved normalized mutual information (NMI) score of 0.626 compared to 0.168 and 0.167, respectively. Furthermore, we found that the centroids can be applied to the segmentation of other slices from the same sample.
* This paper was presented at SPIE Medical Imaging 2018, Houston, TX, USA
Multi-scale Image Fusion Between Pre-operative Clinical CT and X-ray Microtomography of Lung Pathology
Feb 27, 2017
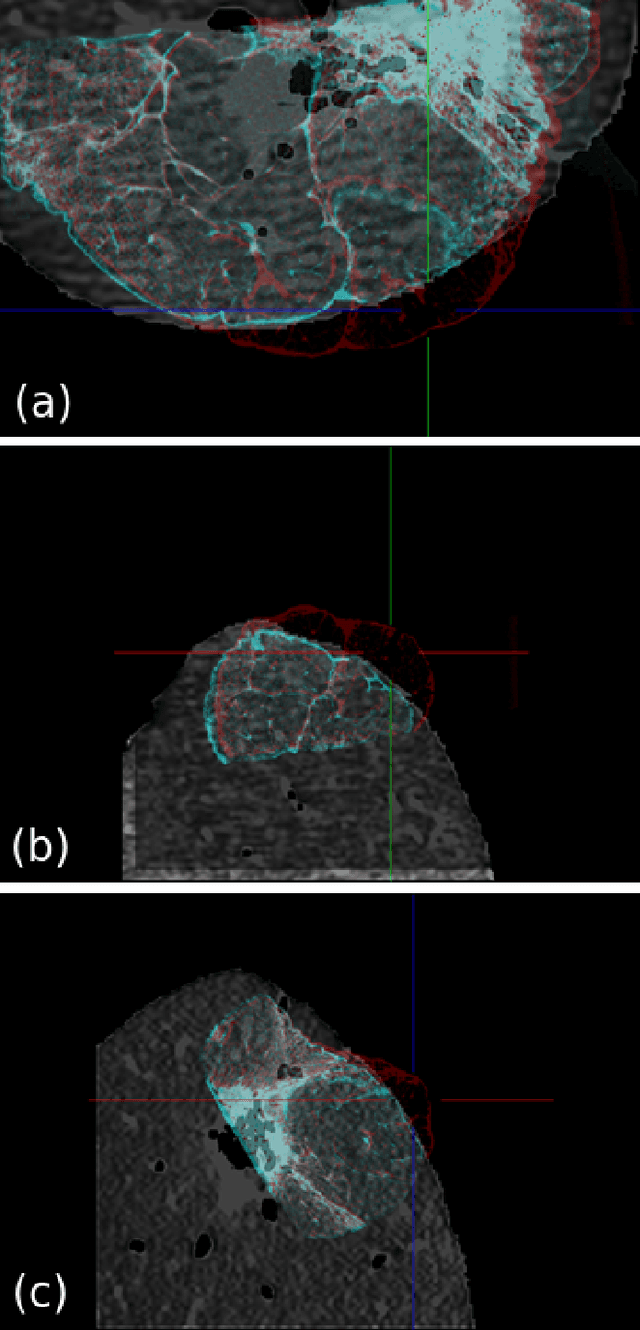
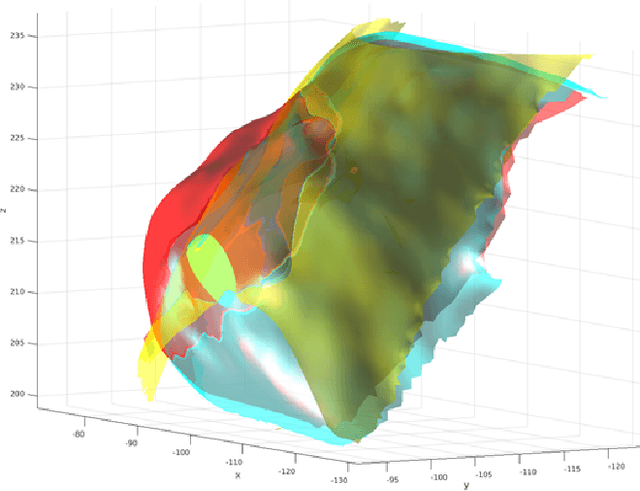

Abstract:Computational anatomy allows the quantitative analysis of organs in medical images. However, most analysis is constrained to the millimeter scale because of the limited resolution of clinical computed tomography (CT). X-ray microtomography ($\mu$CT) on the other hand allows imaging of ex-vivo tissues at a resolution of tens of microns. In this work, we use clinical CT to image lung cancer patients before partial pneumonectomy (resection of pathological lung tissue). The resected specimen is prepared for $\mu$CT imaging at a voxel resolution of 50 $\mu$m (0.05 mm). This high-resolution image of the lung cancer tissue allows further insides into understanding of tumor growth and categorization. For making full use of this additional information, image fusion (registration) needs to be performed in order to re-align the $\mu$CT image with clinical CT. We developed a multi-scale non-rigid registration approach. After manual initialization using a few landmark points and rigid alignment, several levels of non-rigid registration between down-sampled (in the case of $\mu$CT) and up-sampled (in the case of clinical CT) representations of the image are performed. Any non-lung tissue is ignored during the computation of the similarity measure used to guide the registration during optimization. We are able to recover the volume differences introduced by the resection and preparation of the lung specimen. The average ($\pm$ std. dev.) minimum surface distance between $\mu$CT and clinical CT at the resected lung surface is reduced from 3.3 $\pm$ 2.9 (range: [0.1, 15.9]) to 2.3 mm $\pm$ 2.8 (range: [0.0, 15.3]) mm. The alignment of clinical CT with $\mu$CT will allow further registration with even finer resolutions of $\mu$CT (up to 10 $\mu$m resolution) and ultimately with histopathological microscopy images for further macro to micro image fusion that can aid medical image analysis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge