Shekhar Chandra
Structure Guided Manifolds for Discovery of Disease Characteristics
Sep 24, 2022
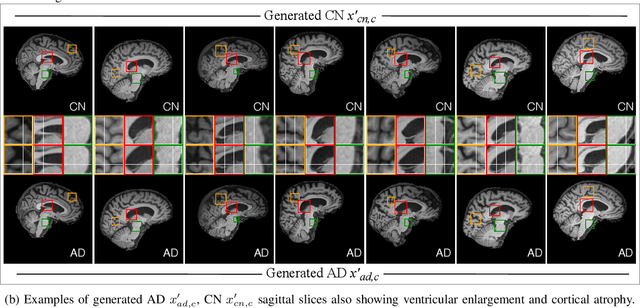
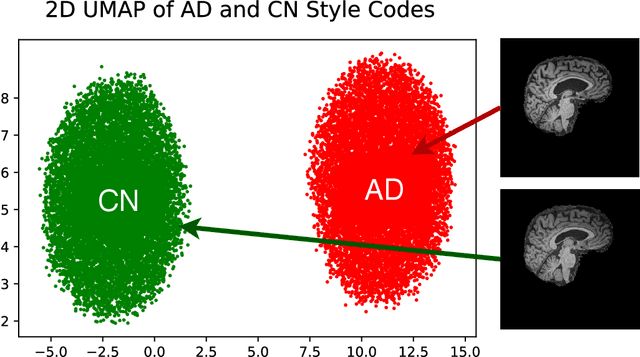

Abstract:In medical image analysis, the subtle visual characteristics of many diseases are challenging to discern, particularly due to the lack of paired data. For example, in mild Alzheimer's Disease (AD), brain tissue atrophy can be difficult to observe from pure imaging data, especially without paired AD and Cognitively Normal ( CN ) data for comparison. This work presents Disease Discovery GAN ( DiDiGAN), a weakly-supervised style-based framework for discovering and visualising subtle disease features. DiDiGAN learns a disease manifold of AD and CN visual characteristics, and the style codes sampled from this manifold are imposed onto an anatomical structural "blueprint" to synthesise paired AD and CN magnetic resonance images (MRIs). To suppress non-disease-related variations between the generated AD and CN pairs, DiDiGAN leverages a structural constraint with cycle consistency and anti-aliasing to enforce anatomical correspondence. When tested on the Alzheimer's Disease Neuroimaging Initiative ( ADNI) dataset, DiDiGAN showed key AD characteristics (reduced hippocampal volume, ventricular enlargement, and atrophy of cortical structures) through synthesising paired AD and CN scans. The qualitative results were backed up by automated brain volume analysis, where systematic pair-wise reductions in brain tissue structures were also measured
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge