Shashata Sawmya
The Birth of Knowledge: Emergent Features across Time, Space, and Scale in Large Language Models
May 26, 2025
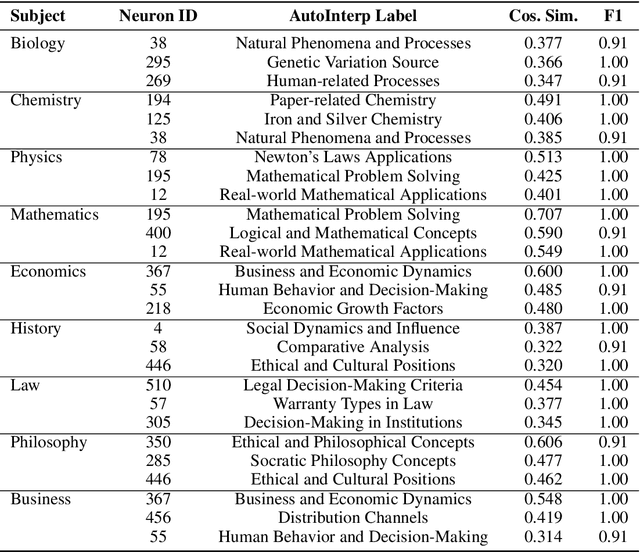

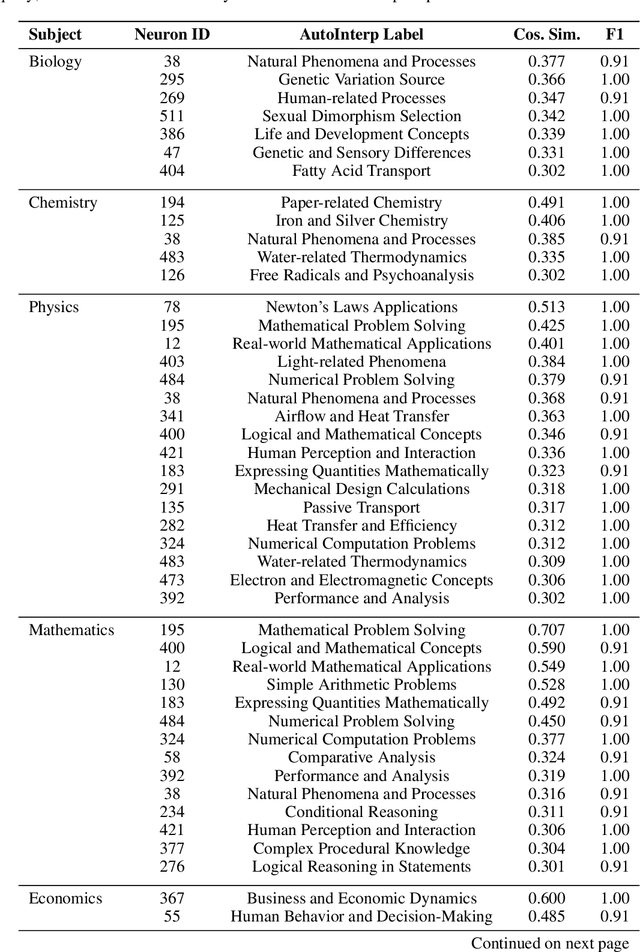
Abstract:This paper studies the emergence of interpretable categorical features within large language models (LLMs), analyzing their behavior across training checkpoints (time), transformer layers (space), and varying model sizes (scale). Using sparse autoencoders for mechanistic interpretability, we identify when and where specific semantic concepts emerge within neural activations. Results indicate clear temporal and scale-specific thresholds for feature emergence across multiple domains. Notably, spatial analysis reveals unexpected semantic reactivation, with early-layer features re-emerging at later layers, challenging standard assumptions about representational dynamics in transformer models.
Cascade Detector Analysis and Application to Biomedical Microscopy
Apr 30, 2025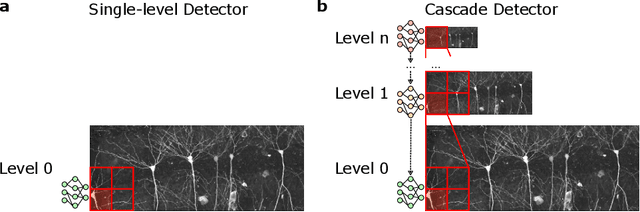
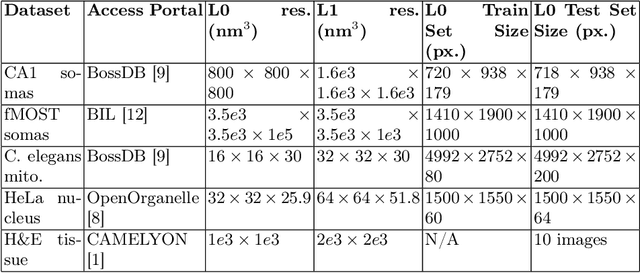
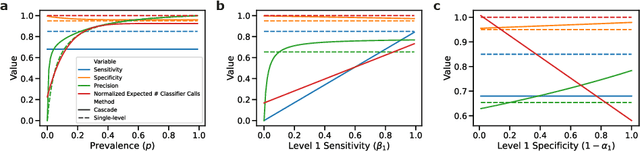

Abstract:As both computer vision models and biomedical datasets grow in size, there is an increasing need for efficient inference algorithms. We utilize cascade detectors to efficiently identify sparse objects in multiresolution images. Given an object's prevalence and a set of detectors at different resolutions with known accuracies, we derive the accuracy, and expected number of classifier calls by a cascade detector. These results generalize across number of dimensions and number of cascade levels. Finally, we compare one- and two-level detectors in fluorescent cell detection, organelle segmentation, and tissue segmentation across various microscopy modalities. We show that the multi-level detector achieves comparable performance in 30-75% less time. Our work is compatible with a variety of computer vision models and data domains.
NeuroADDA: Active Discriminative Domain Adaptation in Connectomic
Mar 08, 2025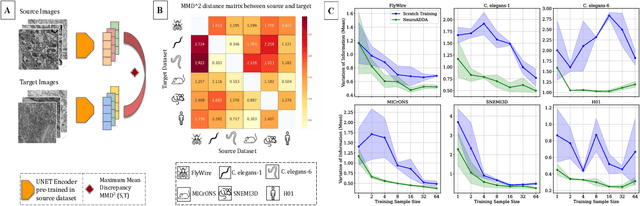
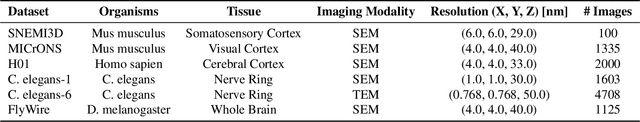
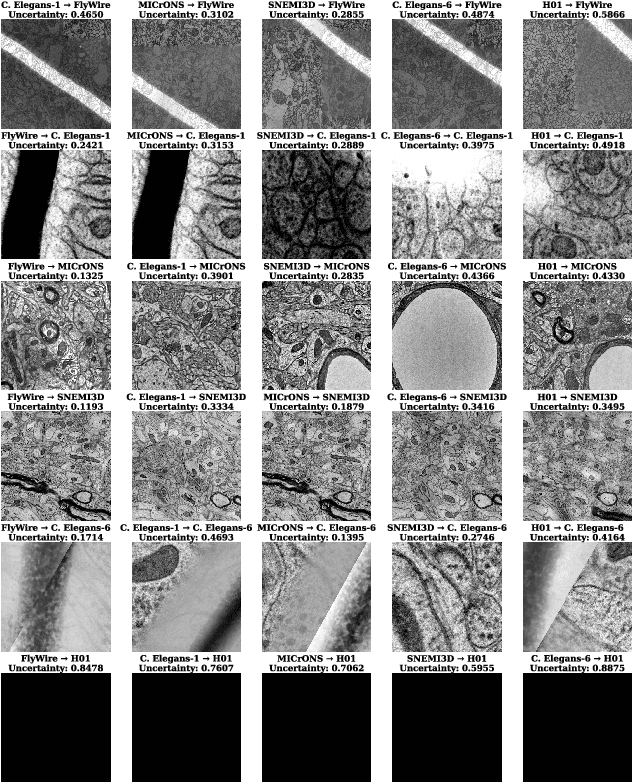


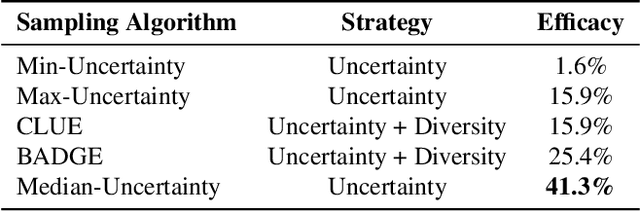
Abstract:Training segmentation models from scratch has been the standard approach for new electron microscopy connectomics datasets. However, leveraging pretrained models from existing datasets could improve efficiency and performance in constrained annotation budget. In this study, we investigate domain adaptation in connectomics by analyzing six major datasets spanning different organisms. We show that, Maximum Mean Discrepancy (MMD) between neuron image distributions serves as a reliable indicator of transferability, and identifies the optimal source domain for transfer learning. Building on this, we introduce NeuroADDA, a method that combines optimal domain selection with source-free active learning to effectively adapt pretrained backbones to a new dataset. NeuroADDA consistently outperforms training from scratch across diverse datasets and fine-tuning sample sizes, with the largest gain observed at $n=4$ samples with a 25-67\% reduction in Variation of Information. Finally, we show that our analysis of distributional differences among neuron images from multiple species in a learned feature space reveals that these domain "distances" correlate with phylogenetic distance among those species.
Sparse Expansion and Neuronal Disentanglement
May 24, 2024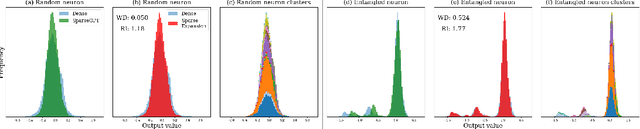
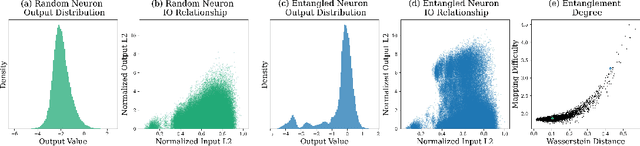
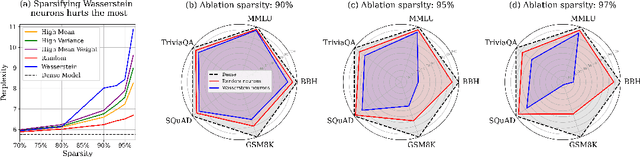
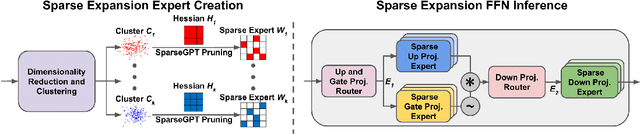
Abstract:We show how to improve the inference efficiency of an LLM by expanding it into a mixture of sparse experts, where each expert is a copy of the original weights, one-shot pruned for a specific cluster of input values. We call this approach $\textit{Sparse Expansion}$. We show that, for models such as Llama 2 70B, as we increase the number of sparse experts, Sparse Expansion outperforms all other one-shot sparsification approaches for the same inference FLOP budget per token, and that this gap grows as sparsity increases, leading to inference speedups. But why? To answer this, we provide strong evidence that the mixture of sparse experts is effectively $\textit{disentangling}$ the input-output relationship of every individual neuron across clusters of inputs. Specifically, sparse experts approximate the dense neuron output distribution with fewer weights by decomposing the distribution into a collection of simpler ones, each with a separate sparse dot product covering it. Interestingly, we show that the Wasserstein distance between a neuron's output distribution and a Gaussian distribution is an indicator of its entanglement level and contribution to the accuracy of the model. Every layer of an LLM has a fraction of highly entangled Wasserstein neurons, and model performance suffers more when these are sparsified as opposed to others.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge