Santa Di Cataldo
Renal Cell Carcinoma subtyping: learning from multi-resolution localization
Nov 14, 2024Abstract:Renal Cell Carcinoma is typically asymptomatic at the early stages for many patients. This leads to a late diagnosis of the tumor, where the curability likelihood is lower, and makes the mortality rate of Renal Cell Carcinoma high, with respect to its incidence rate. To increase the survival chance, a fast and correct categorization of the tumor subtype is paramount. Nowadays, computerized methods, based on artificial intelligence, represent an interesting opportunity to improve the productivity and the objectivity of the microscopy-based Renal Cell Carcinoma diagnosis. Nonetheless, much of their exploitation is hampered by the paucity of annotated dataset, essential for a proficient training of supervised machine learning technologies. This study sets out to investigate a novel self supervised training strategy for machine learning diagnostic tools, based on the multi-resolution nature of the histological samples. We aim at reducing the need of annotated dataset, without significantly reducing the accuracy of the tool. We demonstrate the classification capability of our tool on a whole slide imaging dataset for Renal Cancer subtyping, and we compare our solution with several state-of-the-art classification counterparts.
Neuro-symbolic Empowered Denoising Diffusion Probabilistic Models for Real-time Anomaly Detection in Industry 4.0
Jul 18, 2023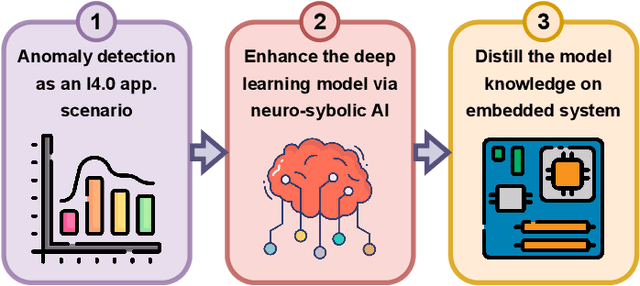
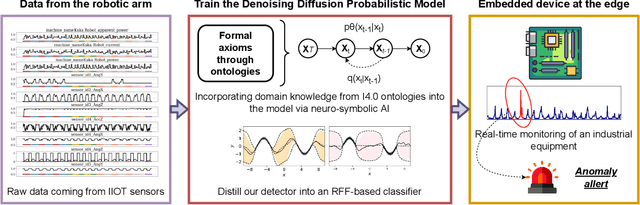
Abstract:Industry 4.0 involves the integration of digital technologies, such as IoT, Big Data, and AI, into manufacturing and industrial processes to increase efficiency and productivity. As these technologies become more interconnected and interdependent, Industry 4.0 systems become more complex, which brings the difficulty of identifying and stopping anomalies that may cause disturbances in the manufacturing process. This paper aims to propose a diffusion-based model for real-time anomaly prediction in Industry 4.0 processes. Using a neuro-symbolic approach, we integrate industrial ontologies in the model, thereby adding formal knowledge on smart manufacturing. Finally, we propose a simple yet effective way of distilling diffusion models through Random Fourier Features for deployment on an embedded system for direct integration into the manufacturing process. To the best of our knowledge, this approach has never been explored before.
Exploiting generative self-supervised learning for the assessment of biological images with lack of annotations: a COVID-19 case-study
Jul 26, 2021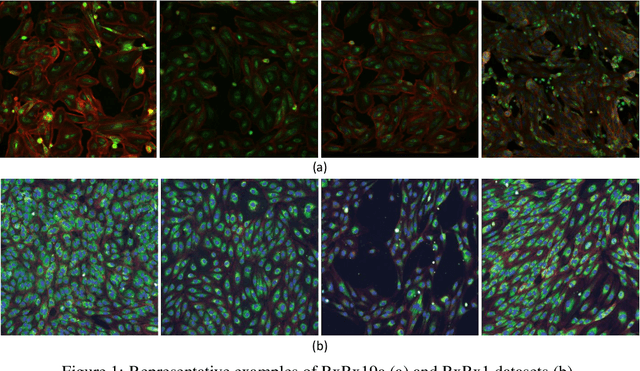
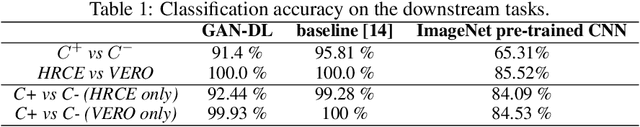

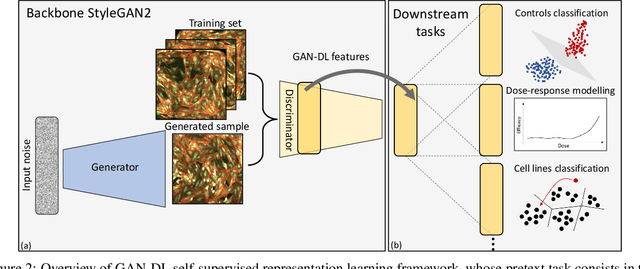
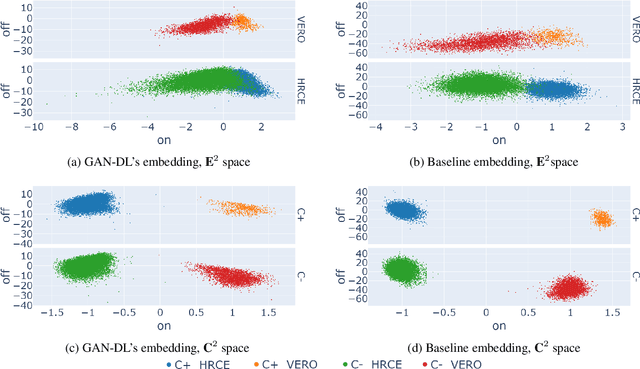
Abstract:Computer-aided analysis of biological images typically requires extensive training on large-scale annotated datasets, which is not viable in many situations. In this paper we present GAN-DL, a Discriminator Learner based on the StyleGAN2 architecture, which we employ for self-supervised image representation learning in the case of fluorescent biological images. We show that Wasserstein Generative Adversarial Networks combined with linear Support Vector Machines enable high-throughput compound screening based on raw images. We demonstrate this by classifying active and inactive compounds tested for the inhibition of SARS-CoV-2 infection in VERO and HRCE cell lines. In contrast to previous methods, our deep learning based approach does not require any annotation besides the one that is normally collected during the sample preparation process. We test our technique on the RxRx19a Sars-CoV-2 image collection. The dataset consists of fluorescent images that were generated to assess the ability of regulatory-approved or in late-stage clinical trials compound to modulate the in vitro infection from SARS-CoV-2 in both VERO and HRCE cell lines. We show that our technique can be exploited not only for classification tasks, but also to effectively derive a dose response curve for the tested treatments, in a self-supervised manner. Lastly, we demonstrate its generalization capabilities by successfully addressing a zero-shot learning task, consisting in the categorization of four different cell types of the RxRx1 fluorescent images collection.
W2WNet: a two-module probabilistic Convolutional Neural Network with embedded data cleansing functionality
Mar 24, 2021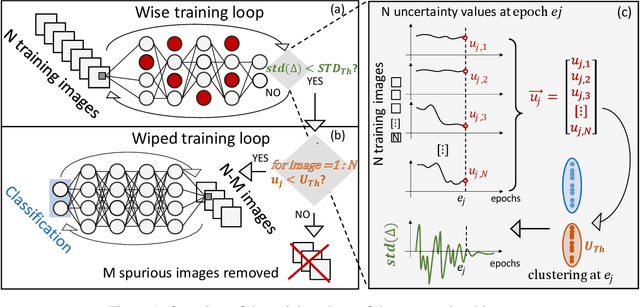
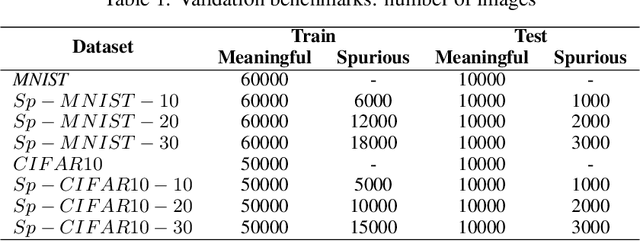
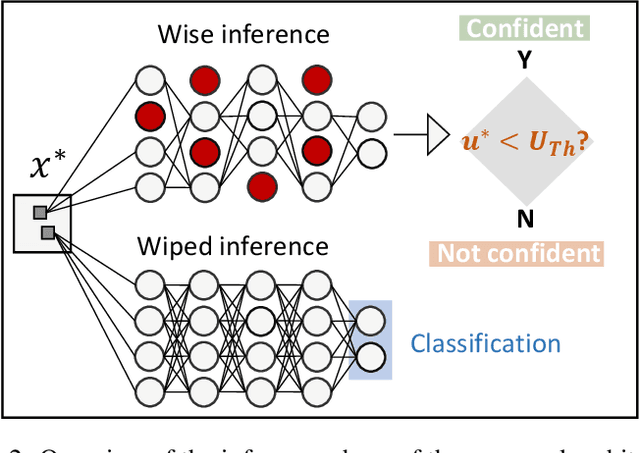
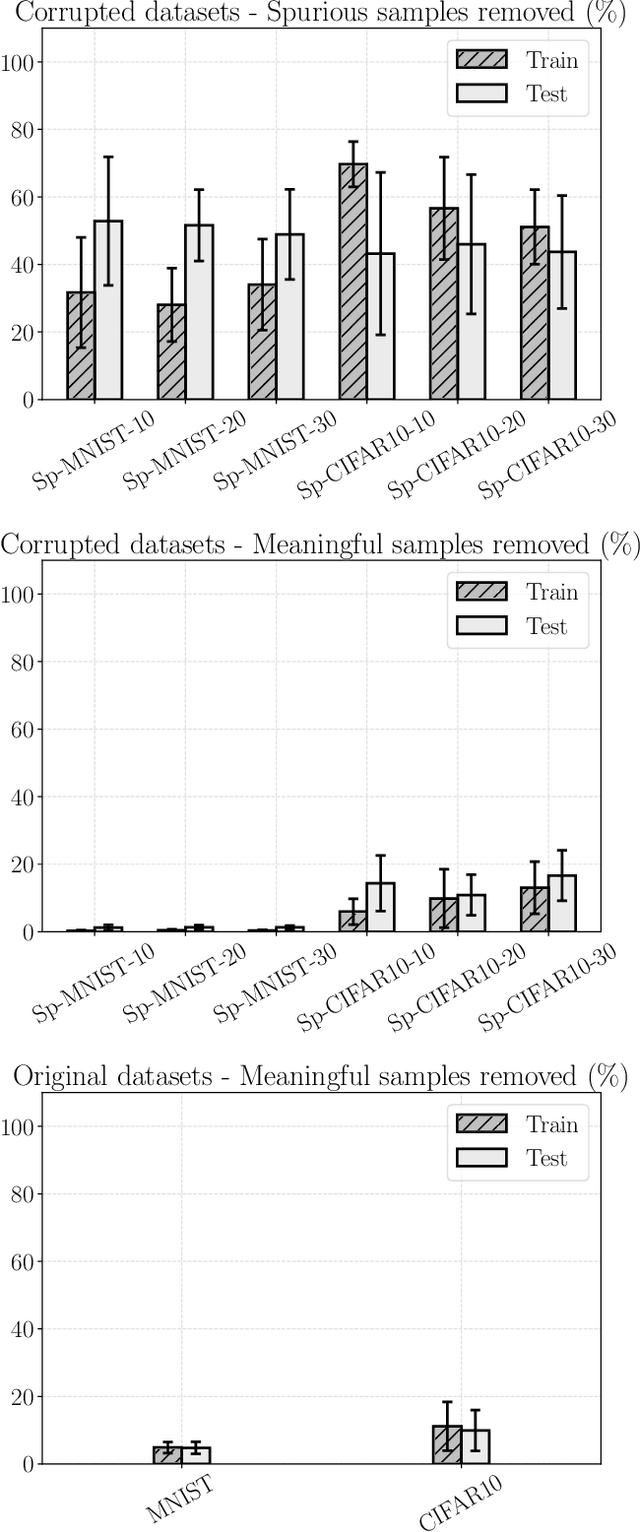
Abstract:Convolutional Neural Networks (CNNs) are supposed to be fed with only high-quality annotated datasets. Nonetheless, in many real-world scenarios, such high quality is very hard to obtain, and datasets may be affected by any sort of image degradation and mislabelling issues. This negatively impacts the performance of standard CNNs, both during the training and the inference phase. To address this issue we propose Wise2WipedNet (W2WNet), a new two-module Convolutional Neural Network, where a Wise module exploits Bayesian inference to identify and discard spurious images during the training, and a Wiped module takes care of the final classification while broadcasting information on the prediction confidence at inference time. The goodness of our solution is demonstrated on a number of public benchmarks addressing different image classification tasks, as well as on a real-world case study on histological image analysis. Overall, our experiments demonstrate that W2WNet is able to identify image degradation and mislabelling issues both at training and at inference time, with a positive impact on the final classification accuracy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge