Sandra E. Black
ROOD-MRI: Benchmarking the robustness of deep learning segmentation models to out-of-distribution and corrupted data in MRI
Mar 11, 2022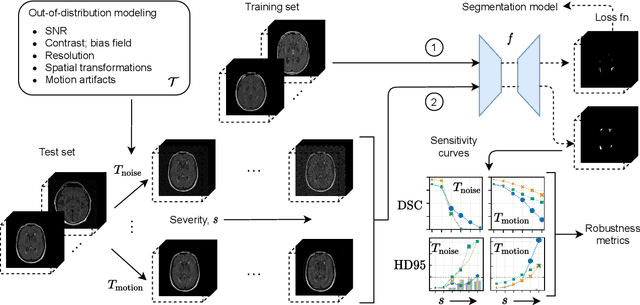
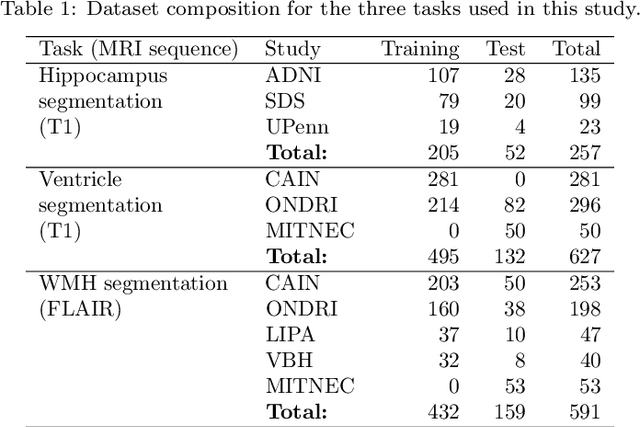
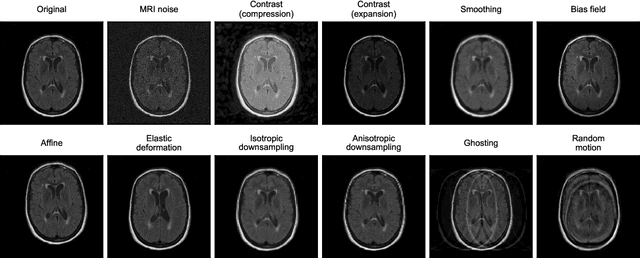
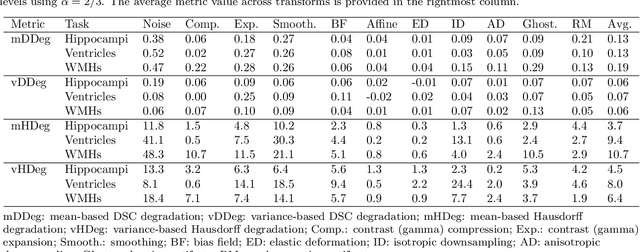
Abstract:Deep artificial neural networks (DNNs) have moved to the forefront of medical image analysis due to their success in classification, segmentation, and detection challenges. A principal challenge in large-scale deployment of DNNs in neuroimage analysis is the potential for shifts in signal-to-noise ratio, contrast, resolution, and presence of artifacts from site to site due to variances in scanners and acquisition protocols. DNNs are famously susceptible to these distribution shifts in computer vision. Currently, there are no benchmarking platforms or frameworks to assess the robustness of new and existing models to specific distribution shifts in MRI, and accessible multi-site benchmarking datasets are still scarce or task-specific. To address these limitations, we propose ROOD-MRI: a platform for benchmarking the Robustness of DNNs to Out-Of-Distribution (OOD) data, corruptions, and artifacts in MRI. The platform provides modules for generating benchmarking datasets using transforms that model distribution shifts in MRI, implementations of newly derived benchmarking metrics for image segmentation, and examples for using the methodology with new models and tasks. We apply our methodology to hippocampus, ventricle, and white matter hyperintensity segmentation in several large studies, providing the hippocampus dataset as a publicly available benchmark. By evaluating modern DNNs on these datasets, we demonstrate that they are highly susceptible to distribution shifts and corruptions in MRI. We show that while data augmentation strategies can substantially improve robustness to OOD data for anatomical segmentation tasks, modern DNNs using augmentation still lack robustness in more challenging lesion-based segmentation tasks. We finally benchmark U-Nets and transformer-based models, finding consistent differences in robustness to particular classes of transforms across architectures.
Perivascular Spaces Segmentation in Brain MRI Using Optimal 3D Filtering
Apr 25, 2017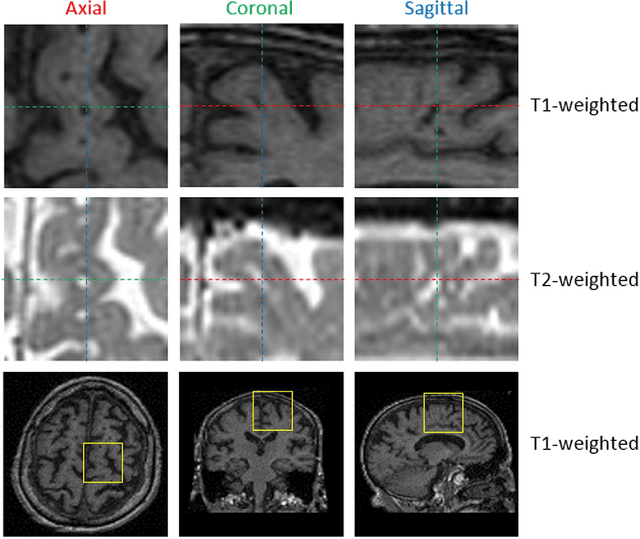
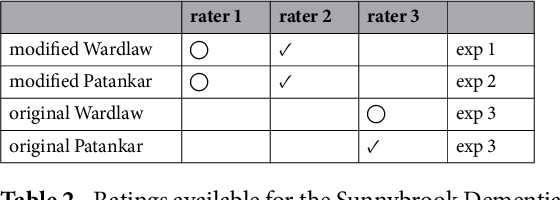
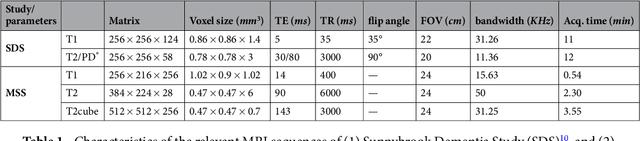
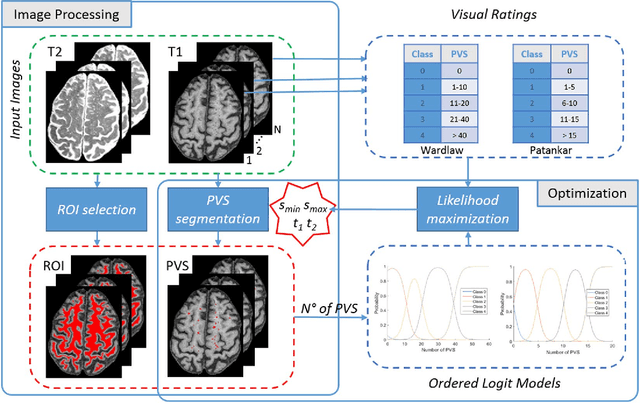

Abstract:Perivascular Spaces (PVS) are a recently recognised feature of Small Vessel Disease (SVD), also indicating neuroinflammation, and are an important part of the brain's circulation and glymphatic drainage system. Quantitative analysis of PVS on Magnetic Resonance Images (MRI) is important for understanding their relationship with neurological diseases. In this work, we propose a segmentation technique based on the 3D Frangi filtering for extraction of PVS from MRI. Based on prior knowledge from neuroradiological ratings of PVS, we used ordered logit models to optimise Frangi filter parameters in response to the variability in the scanner's parameters and study protocols. We optimized and validated our proposed models on two independent cohorts, a dementia sample (N=20) and patients who previously had mild to moderate stroke (N=48). Results demonstrate the robustness and generalisability of our segmentation method. Segmentation-based PVS burden estimates correlated with neuroradiological assessments (Spearman's $\rho$ = 0.74, p $<$ 0.001), suggesting the great potential of our proposed method
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge