Lyndon Boone
A Robust Ensemble Algorithm for Ischemic Stroke Lesion Segmentation: Generalizability and Clinical Utility Beyond the ISLES Challenge
Apr 03, 2024
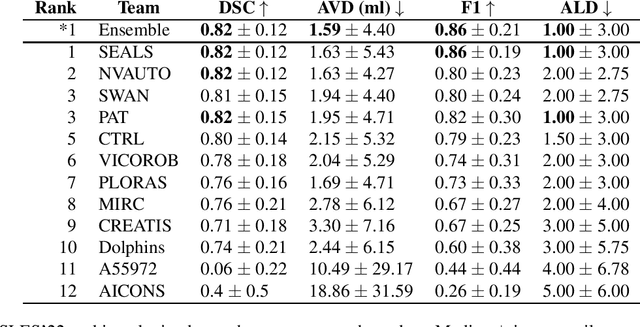
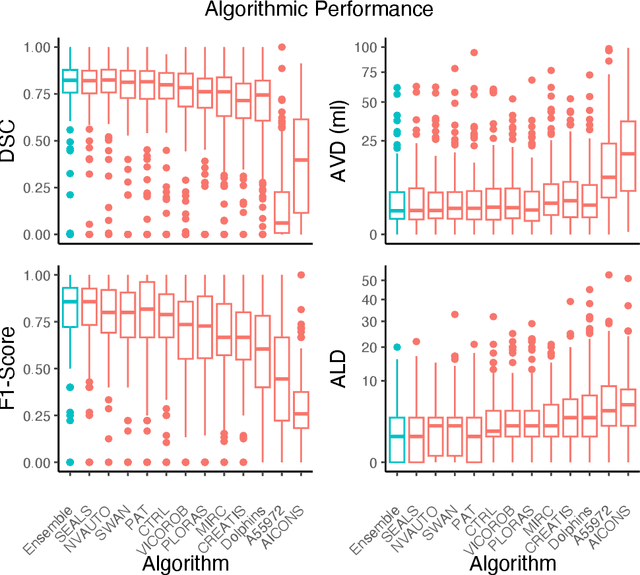
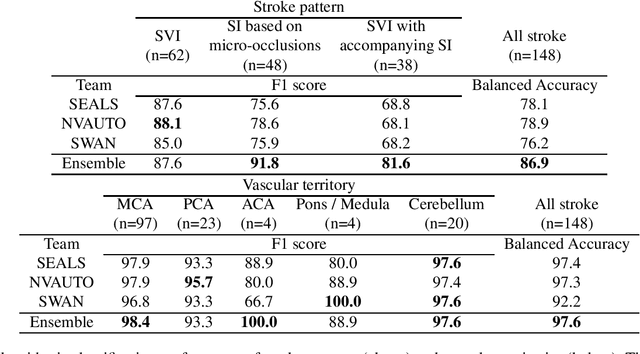
Abstract:Diffusion-weighted MRI (DWI) is essential for stroke diagnosis, treatment decisions, and prognosis. However, image and disease variability hinder the development of generalizable AI algorithms with clinical value. We address this gap by presenting a novel ensemble algorithm derived from the 2022 Ischemic Stroke Lesion Segmentation (ISLES) challenge. ISLES'22 provided 400 patient scans with ischemic stroke from various medical centers, facilitating the development of a wide range of cutting-edge segmentation algorithms by the research community. Through collaboration with leading teams, we combined top-performing algorithms into an ensemble model that overcomes the limitations of individual solutions. Our ensemble model achieved superior ischemic lesion detection and segmentation accuracy on our internal test set compared to individual algorithms. This accuracy generalized well across diverse image and disease variables. Furthermore, the model excelled in extracting clinical biomarkers. Notably, in a Turing-like test, neuroradiologists consistently preferred the algorithm's segmentations over manual expert efforts, highlighting increased comprehensiveness and precision. Validation using a real-world external dataset (N=1686) confirmed the model's generalizability. The algorithm's outputs also demonstrated strong correlations with clinical scores (admission NIHSS and 90-day mRS) on par with or exceeding expert-derived results, underlining its clinical relevance. This study offers two key findings. First, we present an ensemble algorithm (https://github.com/Tabrisrei/ISLES22_Ensemble) that detects and segments ischemic stroke lesions on DWI across diverse scenarios on par with expert (neuro)radiologists. Second, we show the potential for biomedical challenge outputs to extend beyond the challenge's initial objectives, demonstrating their real-world clinical applicability.
ROOD-MRI: Benchmarking the robustness of deep learning segmentation models to out-of-distribution and corrupted data in MRI
Mar 11, 2022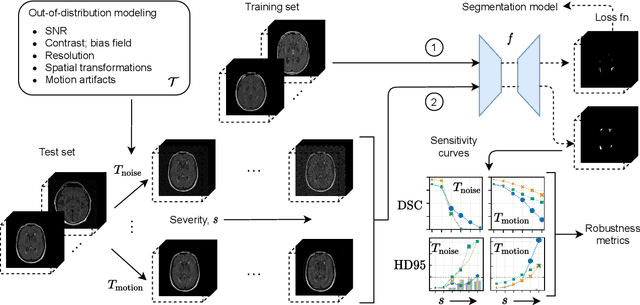
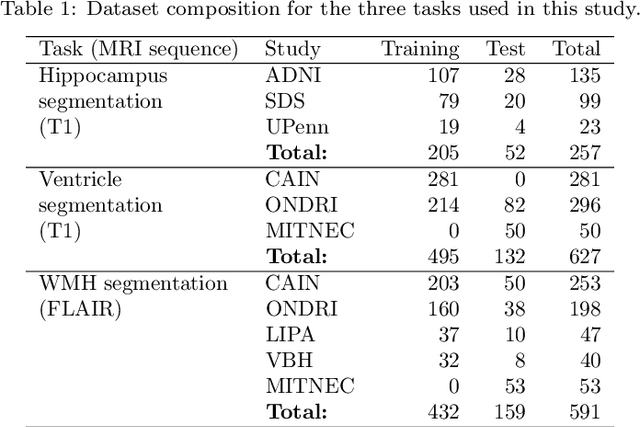
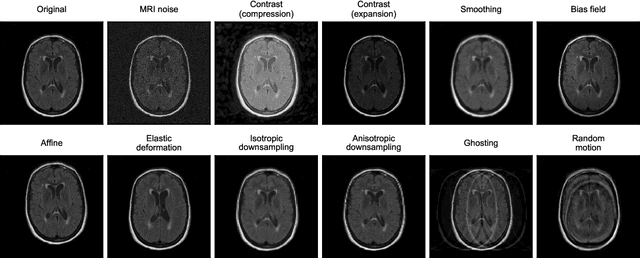
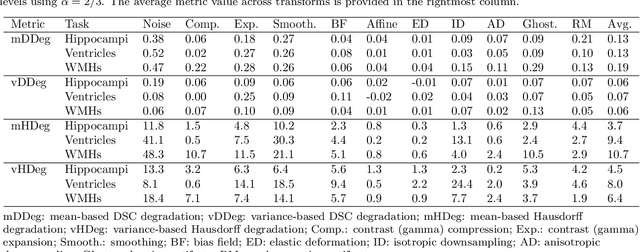
Abstract:Deep artificial neural networks (DNNs) have moved to the forefront of medical image analysis due to their success in classification, segmentation, and detection challenges. A principal challenge in large-scale deployment of DNNs in neuroimage analysis is the potential for shifts in signal-to-noise ratio, contrast, resolution, and presence of artifacts from site to site due to variances in scanners and acquisition protocols. DNNs are famously susceptible to these distribution shifts in computer vision. Currently, there are no benchmarking platforms or frameworks to assess the robustness of new and existing models to specific distribution shifts in MRI, and accessible multi-site benchmarking datasets are still scarce or task-specific. To address these limitations, we propose ROOD-MRI: a platform for benchmarking the Robustness of DNNs to Out-Of-Distribution (OOD) data, corruptions, and artifacts in MRI. The platform provides modules for generating benchmarking datasets using transforms that model distribution shifts in MRI, implementations of newly derived benchmarking metrics for image segmentation, and examples for using the methodology with new models and tasks. We apply our methodology to hippocampus, ventricle, and white matter hyperintensity segmentation in several large studies, providing the hippocampus dataset as a publicly available benchmark. By evaluating modern DNNs on these datasets, we demonstrate that they are highly susceptible to distribution shifts and corruptions in MRI. We show that while data augmentation strategies can substantially improve robustness to OOD data for anatomical segmentation tasks, modern DNNs using augmentation still lack robustness in more challenging lesion-based segmentation tasks. We finally benchmark U-Nets and transformer-based models, finding consistent differences in robustness to particular classes of transforms across architectures.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge