Rolf F Schulte
Pinv-Recon: Generalized MR Image Reconstruction via Pseudoinversion of the Encoding Matrix
Oct 08, 2024
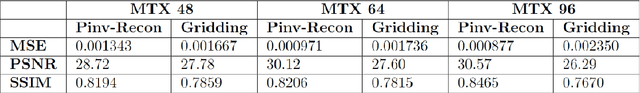
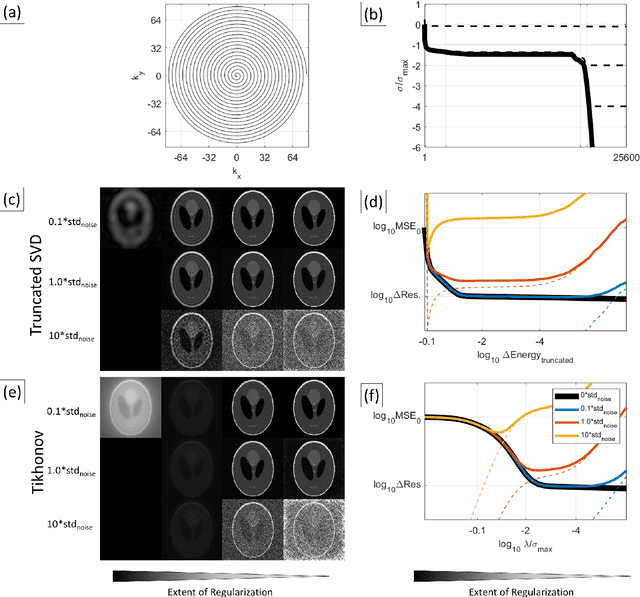
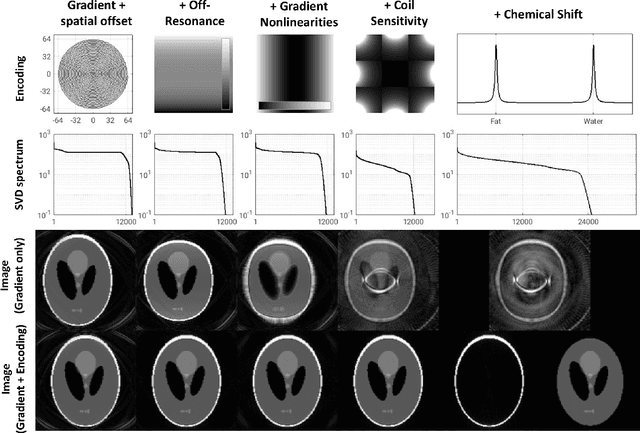
Abstract:Purpose: To present a novel generalized MR image reconstruction based on pseudoinversion of the encoding matrix (Pinv-Recon) as a simple yet powerful method, and demonstrate its computational feasibility for diverse MR imaging applications. Methods: MR image encoding constitutes a linear mapping of the unknown image to the measured k-space data mediated via an encoding matrix ($ data = Encode \times image$). Pinv-Recon addresses MR image reconstruction as a linear inverse problem ($image = Encode^{-1} \times data$), explicitly calculating the Moore-Penrose pseudoinverse of the encoding matrix using truncated singular value decomposition (tSVD). Using a discretized, algebraic notation, we demonstrate constructing a generalized encoding matrix by stacking relevant encoding mechanisms (e.g., gradient encoding, coil sensitivity encoding, chemical shift inversion) and encoding distortions (e.g., off-center positioning, B$_0$ inhomogeneity, spatiotemporal gradient imperfections, transient relaxation effects). Iterative reconstructions using the explicit generalized encoding matrix, and the computation of the spatial-response-function (SRF) and noise amplification, were demonstrated. Results: We evaluated the computation times and memory requirements (time ~ (size of the encoding matrix)$^{1.4}$). Using the Shepp-Logan phantom, we demonstrated the versatility of the method for various intertwined MR image encoding and distortion mechanisms, achieving better MSE, PSNR and SSIM metrics than conventional methods. A diversity of datasets, including the ISMRM CG-SENSE challenge, were used to validate Pinv-Recon. Conclusion: Although pseudo-inversion of large encoding matrices was once deemed computationally intractable, recent advances make Pinv-Recon feasible. It has great promise for both research and clinical applications, and for educational use.
Joint alignment and reconstruction of multislice dynamic MRI using variational manifold learning
Nov 21, 2021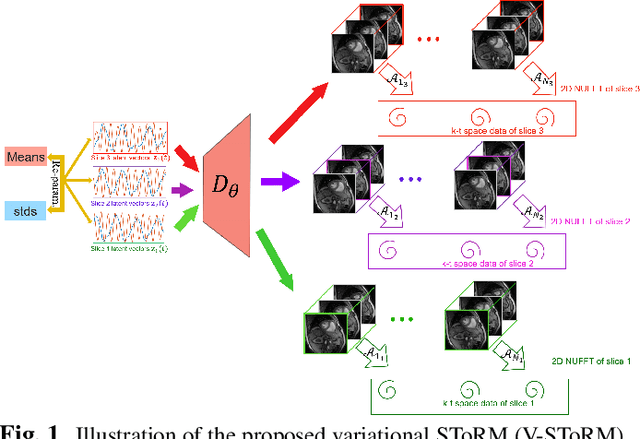
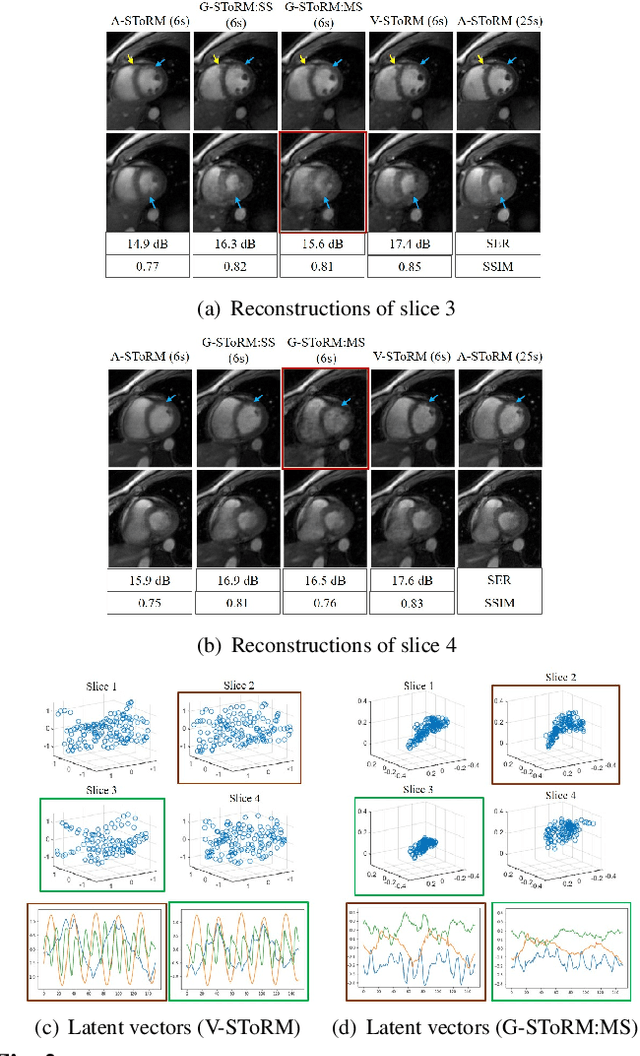

Abstract:Free-breathing cardiac MRI schemes are emerging as competitive alternatives to breath-held cine MRI protocols, enabling applicability to pediatric and other population groups that cannot hold their breath. Because the data from the slices are acquired sequentially, the cardiac/respiratory motion patterns may be different for each slice; current free-breathing approaches perform independent recovery of each slice. In addition to not being able to exploit the inter-slice redundancies, manual intervention or sophisticated post-processing methods are needed to align the images post-recovery for quantification. To overcome these challenges, we propose an unsupervised variational deep manifold learning scheme for the joint alignment and reconstruction of multislice dynamic MRI. The proposed scheme jointly learns the parameters of the deep network as well as the latent vectors for each slice, which capture the motion-induced dynamic variations, from the k-t space data of the specific subject. The variational framework minimizes the non-uniqueness in the representation, thus offering improved alignment and reconstructions.
Qunatification of Metabolites in MR Spectroscopic Imaging using Machine Learning
May 25, 2018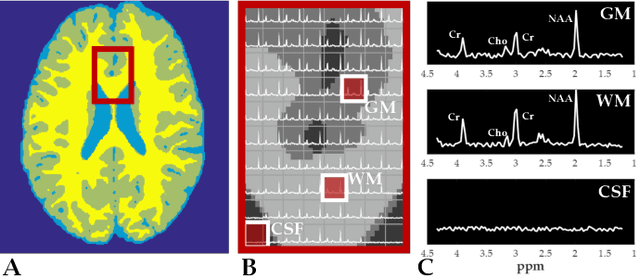
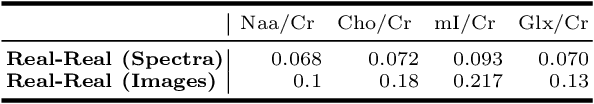
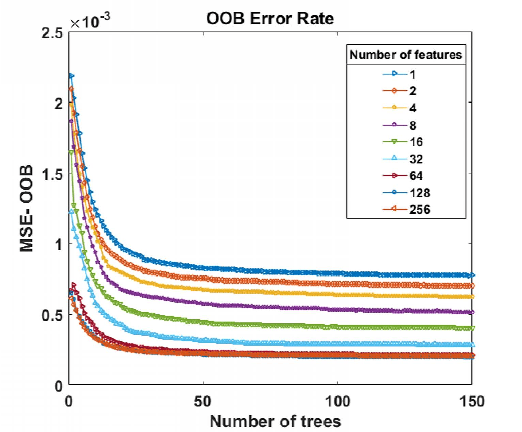
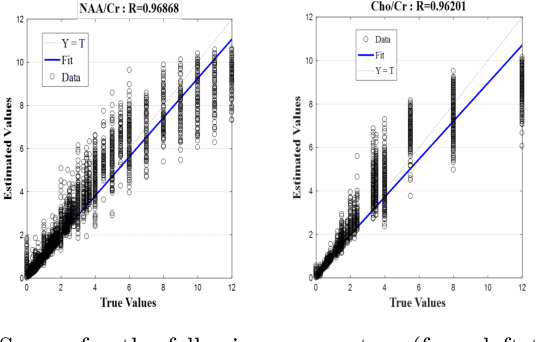
Abstract:Magnetic Resonance Spectroscopic Imaging (MRSI) is a clinical imaging modality for measuring tissue metabolite levels in-vivo. An accurate estimation of spectral parameters allows for better assessment of spectral quality and metabolite concentration levels. The current gold standard quantification method is the LCModel - a commercial fitting tool. However, this fails for spectra having poor signal-to-noise ratio (SNR) or a large number of artifacts. This paper introduces a framework based on random forest regression for accurate estimation of the output parameters of a model based analysis of MR spectroscopy data. The goal of our proposed framework is to learn the spectral features from a training set comprising of different variations of both simulated and in-vivo brain spectra and then use this learning for the subsequent metabolite quantification. Experiments involve training and testing on simulated and in-vivo human brain spectra. We estimate parameters such as concentration of metabolites and compare our results with that from the LCModel.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge