Monique M. B Breteler
Automated Olfactory Bulb Segmentation on High Resolutional T2-Weighted MRI
Aug 09, 2021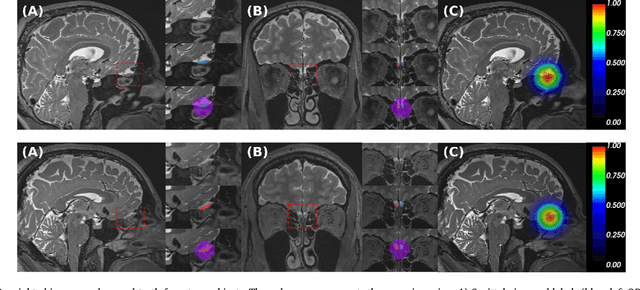
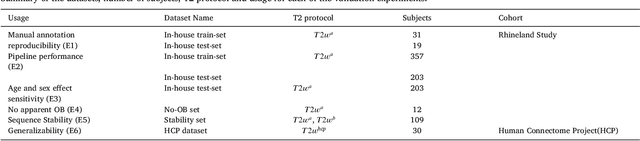

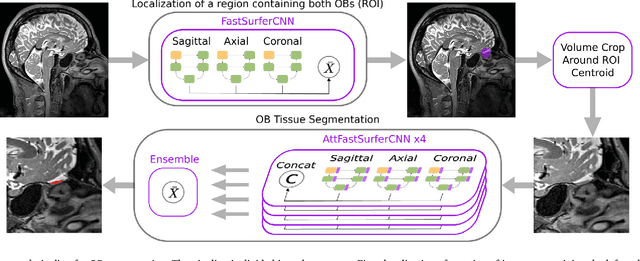
Abstract:The neuroimage analysis community has neglected the automated segmentation of the olfactory bulb (OB) despite its crucial role in olfactory function. The lack of an automatic processing method for the OB can be explained by its challenging properties. Nonetheless, recent advances in MRI acquisition techniques and resolution have allowed raters to generate more reliable manual annotations. Furthermore, the high accuracy of deep learning methods for solving semantic segmentation problems provides us with an option to reliably assess even small structures. In this work, we introduce a novel, fast, and fully automated deep learning pipeline to accurately segment OB tissue on sub-millimeter T2-weighted (T2w) whole-brain MR images. To this end, we designed a three-stage pipeline: (1) Localization of a region containing both OBs using FastSurferCNN, (2) Segmentation of OB tissue within the localized region through four independent AttFastSurferCNN - a novel deep learning architecture with a self-attention mechanism to improve modeling of contextual information, and (3) Ensemble of the predicted label maps. The OB pipeline exhibits high performance in terms of boundary delineation, OB localization, and volume estimation across a wide range of ages in 203 participants of the Rhineland Study. Moreover, it also generalizes to scans of an independent dataset never encountered during training, the Human Connectome Project (HCP), with different acquisition parameters and demographics, evaluated in 30 cases at the native 0.7mm HCP resolution, and the default 0.8mm pipeline resolution. We extensively validated our pipeline not only with respect to segmentation accuracy but also to known OB volume effects, where it can sensitively replicate age effects.
FatSegNet : A Fully Automated Deep Learning Pipeline for Adipose Tissue Segmentation on Abdominal Dixon MRI
Apr 03, 2019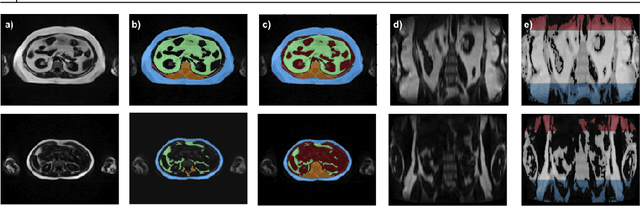

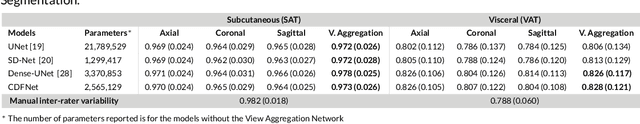
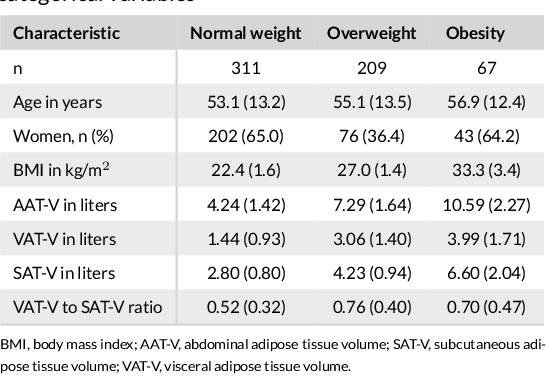
Abstract:Purpose: Development of a fast and fully automated deep learning pipeline (FatSegNet) to accurately identify, segment, and quantify abdominal adipose tissue on Dixon MRI from the Rhineland Study - a large prospective population-based study. Method: FatSegNet is composed of three stages: (i) consistent localization of the abdominal region using two 2D-Competitive Dense Fully Convolutional Networks (CDFNet), (ii) segmentation of adipose tissue on three views by independent CDFNets, and (iii) view aggregation. FatSegNet is trained with 33 manually annotated subjects, and validated by: 1) comparison of segmentation accuracy against a testingset covering a wide range of body mass index (BMI), 2) test-retest reliability, and 3) robustness in a large cohort study. Results: The CDFNet demonstrates increased robustness compared to traditional deep learning networks. FatSegNet dice score outperforms manual raters on the abdominal visceral adipose tissue (VAT, 0.828 vs. 0.788), and produces comparable results on subcutaneous adipose tissue (SAT, 0.973 vs. 0.982). The pipeline has very small test-retest absolute percentage difference and excellent agreement between scan sessions (VAT: APD = 2.957%, ICC=0.998 and SAT: APD= 3.254%, ICC=0.996). Conclusion: FatSegNet can reliably analyze a 3D Dixon MRI in1 min. It generalizes well to different body shapes, sensitively replicates known VAT and SAT volume effects in a large cohort study, and permits localized analysis of fat compartments.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge