Mohiuddin Ahmad
Acute Lymphoblastic Leukemia Detection from Microscopic Images Using Weighted Ensemble of Convolutional Neural Networks
May 09, 2021
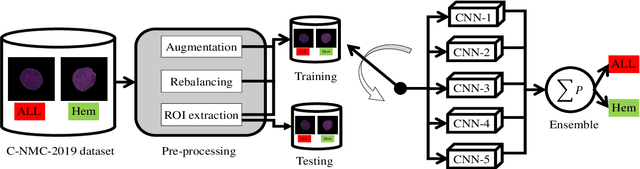
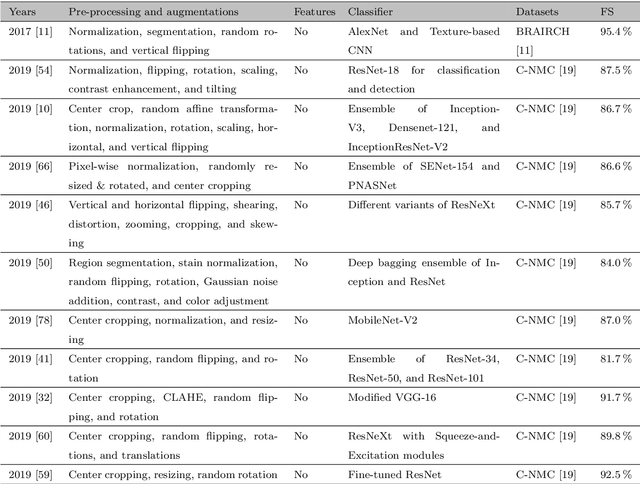
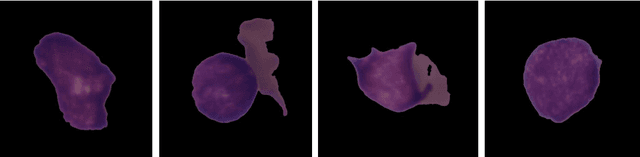
Abstract:Acute Lymphoblastic Leukemia (ALL) is a blood cell cancer characterized by numerous immature lymphocytes. Even though automation in ALL prognosis is an essential aspect of cancer diagnosis, it is challenging due to the morphological correlation between malignant and normal cells. The traditional ALL classification strategy demands experienced pathologists to carefully read the cell images, which is arduous, time-consuming, and often suffers inter-observer variations. This article has automated the ALL detection task from microscopic cell images, employing deep Convolutional Neural Networks (CNNs). We explore the weighted ensemble of different deep CNNs to recommend a better ALL cell classifier. The weights for the ensemble candidate models are estimated from their corresponding metrics, such as accuracy, F1-score, AUC, and kappa values. Various data augmentations and pre-processing are incorporated for achieving a better generalization of the network. We utilize the publicly available C-NMC-2019 ALL dataset to conduct all the comprehensive experiments. Our proposed weighted ensemble model, using the kappa values of the ensemble candidates as their weights, has outputted a weighted F1-score of 88.6 %, a balanced accuracy of 86.2 %, and an AUC of 0.941 in the preliminary test set. The qualitative results displaying the gradient class activation maps confirm that the introduced model has a concentrated learned region. In contrast, the ensemble candidate models, such as Xception, VGG-16, DenseNet-121, MobileNet, and InceptionResNet-V2, separately produce coarse and scatter learned areas for most example cases. Since the proposed kappa value-based weighted ensemble yields a better result for the aimed task in this article, it can experiment in other domains of medical diagnostic applications.
Dermo-DOCTOR: A framework for concurrent skin lesion detection and recognition using a deep convolutional neural network with end-to-end dual encoders
Feb 23, 2021
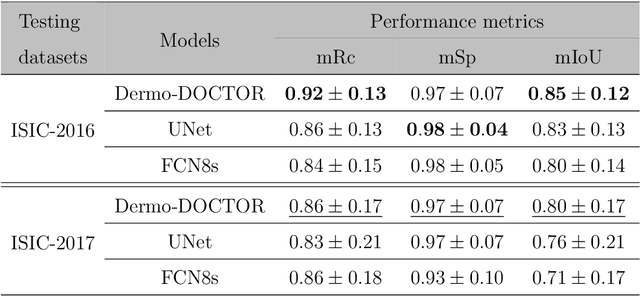
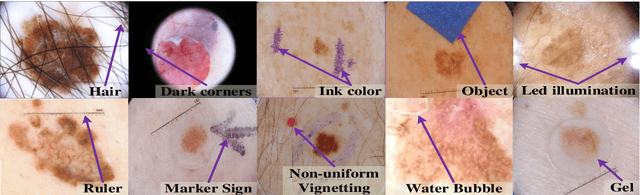
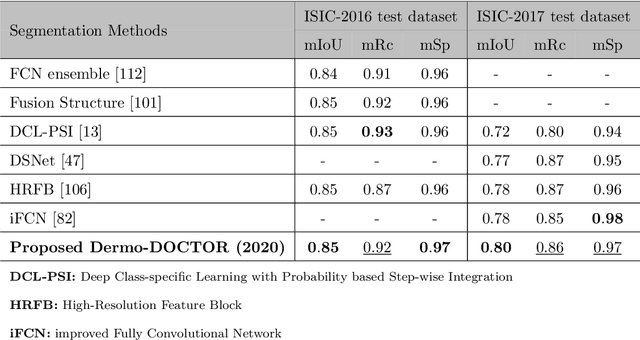
Abstract:Automated skin lesion analysis for simultaneous detection and recognition is still challenging for inter-class homogeneity and intra-class heterogeneity, leading to low generic capability of a Single Convolutional Neural Network (CNN) with limited datasets. This article proposes an end-to-end deep CNN-based framework for simultaneous detection and recognition of the skin lesions, named Dermo-DOCTOR, consisting of two encoders. The feature maps from two encoders are fused channel-wise, called Fused Feature Map (FFM). The FFM is utilized for decoding in the detection sub-network, concatenating each stage of two encoders' outputs with corresponding decoder layers to retrieve the lost spatial information due to pooling in the encoders. For the recognition sub-network, the outputs of three fully connected layers, utilizing feature maps of two encoders and FFM, are aggregated to obtain a final lesion class. We train and evaluate the proposed Dermo-Doctor utilizing two publicly available benchmark datasets, such as ISIC-2016 and ISIC-2017. The achieved segmentation results exhibit mean intersection over unions of 85.0 % and 80.0 % respectively for ISIC-2016 and ISIC-2017 test datasets. The proposed Dermo-DOCTOR also demonstrates praiseworthy success in lesion recognition, providing the areas under the receiver operating characteristic curves of 0.98 and 0.91 respectively for those two datasets. The experimental results show that the proposed Dermo-DOCTOR outperforms the alternative methods mentioned in the literature, designed for skin lesion detection and recognition. As the Dermo-DOCTOR provides better-results on two different test datasets, even with limited training data, it can be an auspicious computer-aided assistive tool for dermatologists.
Multi-class probabilistic atlas-based whole heart segmentation method in cardiac CT and MRI
Feb 03, 2021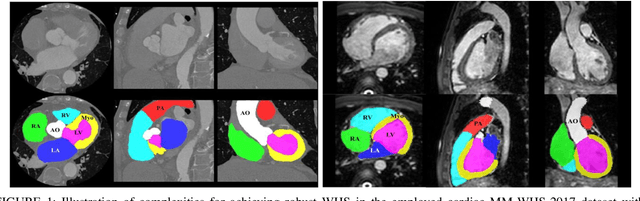
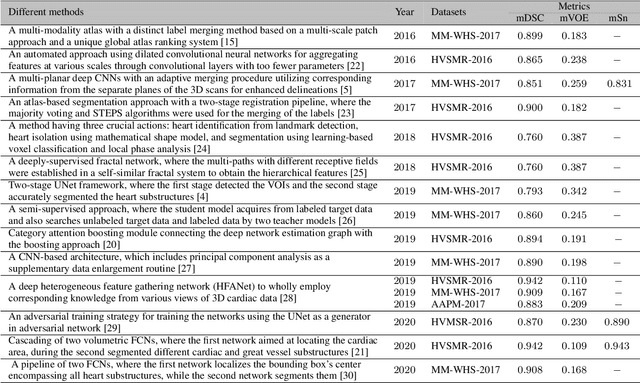
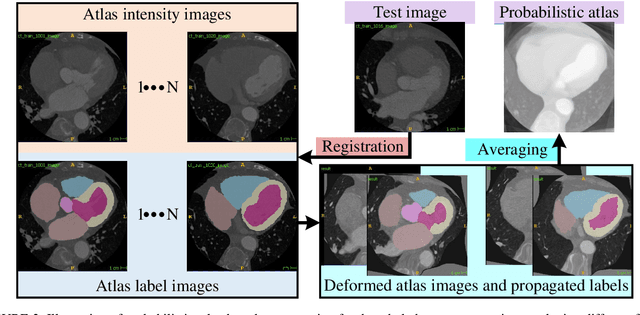
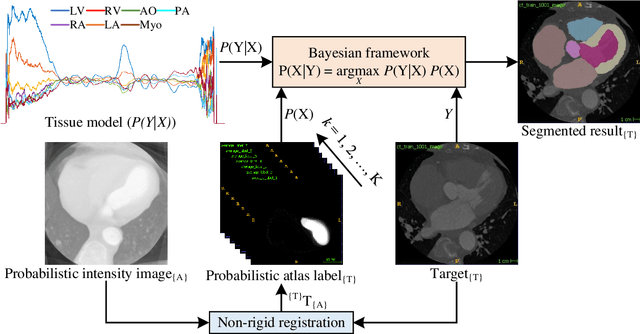
Abstract:Accurate and robust whole heart substructure segmentation is crucial in developing clinical applications, such as computer-aided diagnosis and computer-aided surgery. However, segmentation of different heart substructures is challenging because of inadequate edge or boundary information, the complexity of the background and texture, and the diversity in different substructures' sizes and shapes. This article proposes a framework for multi-class whole heart segmentation employing non-rigid registration-based probabilistic atlas incorporating the Bayesian framework. We also propose a non-rigid registration pipeline utilizing a multi-resolution strategy for obtaining the highest attainable mutual information between the moving and fixed images. We further incorporate non-rigid registration into the expectation-maximization algorithm and implement different deep convolutional neural network-based encoder-decoder networks for ablation studies. All the extensive experiments are conducted utilizing the publicly available dataset for the whole heart segmentation containing 20 MRI and 20 CT cardiac images. The proposed approach exhibits an encouraging achievement, yielding a mean volume overlapping error of 14.5 % for CT scans exceeding the state-of-the-art results by a margin of 1.3 % in terms of the same metric. As the proposed approach provides better-results to delineate the different substructures of the heart, it can be a medical diagnostic aiding tool for helping experts with quicker and more accurate results.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge