Michał Januszewski
Discovering Mechanistic Models of Neural Activity: System Identification in an in Silico Zebrafish
Feb 04, 2026Abstract:Constructing mechanistic models of neural circuits is a fundamental goal of neuroscience, yet verifying such models is limited by the lack of ground truth. To rigorously test model discovery, we establish an in silico testbed using neuromechanical simulations of a larval zebrafish as a transparent ground truth. We find that LLM-based tree search autonomously discovers predictive models that significantly outperform established forecasting baselines. Conditioning on sensory drive is necessary but not sufficient for faithful system identification, as models exploit statistical shortcuts. Structural priors prove essential for enabling robust out-of-distribution generalization and recovery of interpretable mechanistic models. Our insights provide guidance for modeling real-world neural recordings and offer a broader template for AI-driven scientific discovery.
ZAPBench: A Benchmark for Whole-Brain Activity Prediction in Zebrafish
Mar 04, 2025



Abstract:Data-driven benchmarks have led to significant progress in key scientific modeling domains including weather and structural biology. Here, we introduce the Zebrafish Activity Prediction Benchmark (ZAPBench) to measure progress on the problem of predicting cellular-resolution neural activity throughout an entire vertebrate brain. The benchmark is based on a novel dataset containing 4d light-sheet microscopy recordings of over 70,000 neurons in a larval zebrafish brain, along with motion stabilized and voxel-level cell segmentations of these data that facilitate development of a variety of forecasting methods. Initial results from a selection of time series and volumetric video modeling approaches achieve better performance than naive baseline methods, but also show room for further improvement. The specific brain used in the activity recording is also undergoing synaptic-level anatomical mapping, which will enable future integration of detailed structural information into forecasting methods.
Flood-Filling Networks
Nov 01, 2016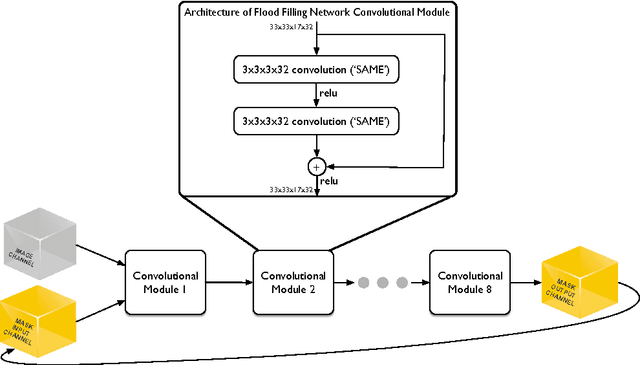
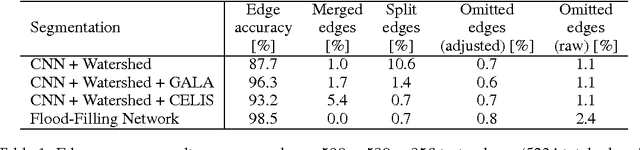
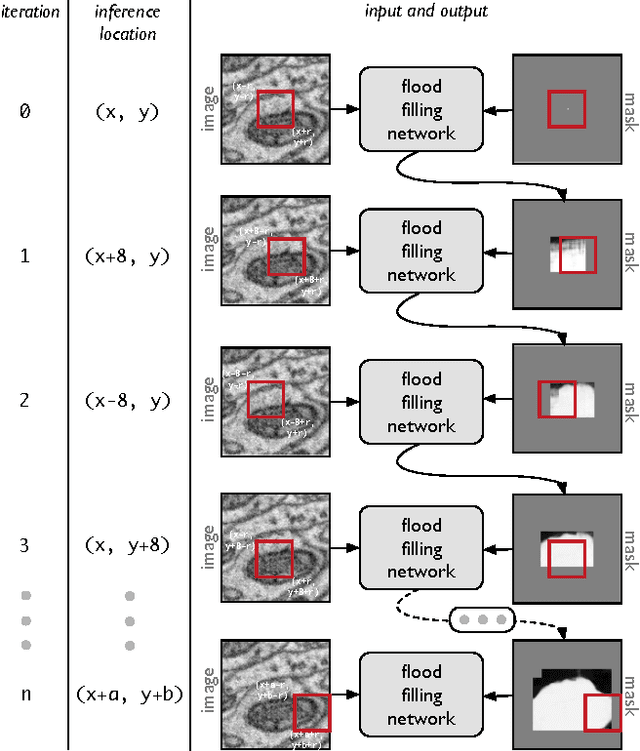
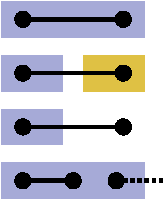
Abstract:State-of-the-art image segmentation algorithms generally consist of at least two successive and distinct computations: a boundary detection process that uses local image information to classify image locations as boundaries between objects, followed by a pixel grouping step such as watershed or connected components that clusters pixels into segments. Prior work has varied the complexity and approach employed in these two steps, including the incorporation of multi-layer neural networks to perform boundary prediction, and the use of global optimizations during pixel clustering. We propose a unified and end-to-end trainable machine learning approach, flood-filling networks, in which a recurrent 3d convolutional network directly produces individual segments from a raw image. The proposed approach robustly segments images with an unknown and variable number of objects as well as highly variable object sizes. We demonstrate the approach on a challenging 3d image segmentation task, connectomic reconstruction from volume electron microscopy data, on which flood-filling neural networks substantially improve accuracy over other state-of-the-art methods. The proposed approach can replace complex multi-step segmentation pipelines with a single neural network that is learned end-to-end.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge