Mary F. Paine
Developing a Knowledge Graph Framework for Pharmacokinetic Natural Product-Drug Interactions
Sep 24, 2022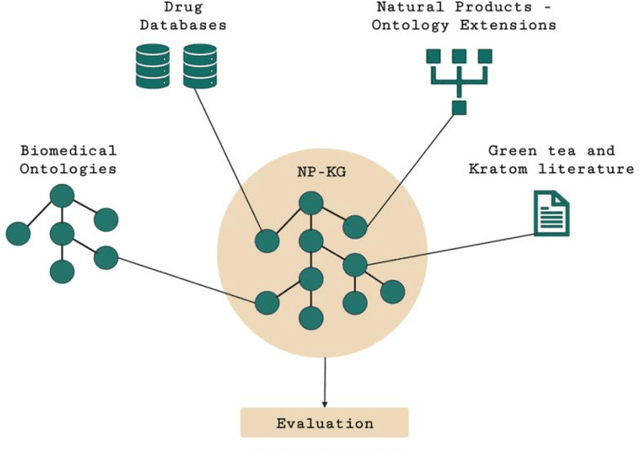
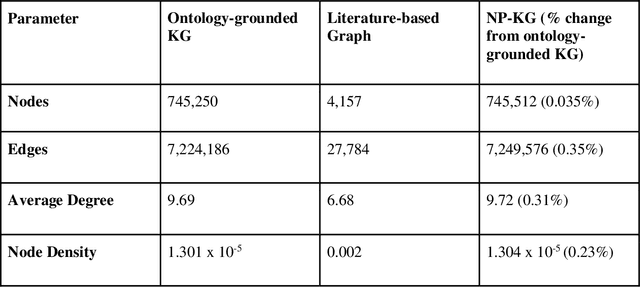
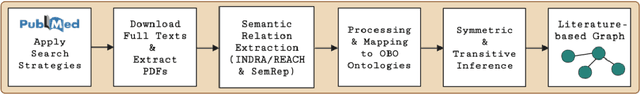
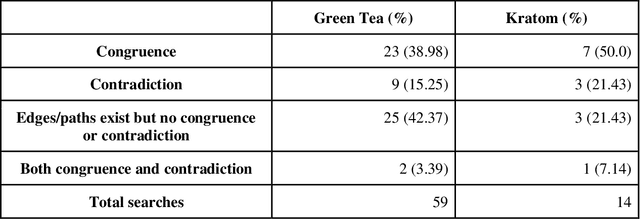
Abstract:Pharmacokinetic natural product-drug interactions (NPDIs) occur when botanical natural products are co-consumed with pharmaceutical drugs. Understanding mechanisms of NPDIs is key to preventing adverse events. We constructed a knowledge graph framework, NP-KG, as a step toward computational discovery of pharmacokinetic NPDIs. NP-KG is a heterogeneous KG with biomedical ontologies, linked data, and full texts of the scientific literature, constructed with the Phenotype Knowledge Translator framework and the semantic relation extraction systems, SemRep and Integrated Network and Dynamic Reasoning Assembler. NP-KG was evaluated with case studies of pharmacokinetic green tea- and kratom-drug interactions through path searches and meta-path discovery to determine congruent and contradictory information compared to ground truth data. The fully integrated NP-KG consisted of 745,512 nodes and 7,249,576 edges. Evaluation of NP-KG resulted in congruent (38.98% for green tea, 50% for kratom), contradictory (15.25% for green tea, 21.43% for kratom), and both congruent and contradictory (15.25% for green tea, 21.43% for kratom) information. Potential pharmacokinetic mechanisms for several purported NPDIs, including the green tea-raloxifene, green tea-nadolol, kratom-midazolam, kratom-quetiapine, and kratom-venlafaxine interactions were congruent with the published literature. NP-KG is the first KG to integrate biomedical ontologies with full texts of the scientific literature focused on natural products. We demonstrate the application of NP-KG to identify pharmacokinetic interactions involving enzymes, transporters, and pharmaceutical drugs. We envision that NP-KG will facilitate improved human-machine collaboration to guide researchers in future studies of pharmacokinetic NPDIs. The NP-KG framework is publicly available at https://doi.org/10.5281/zenodo.6814507 and https://github.com/sanyabt/np-kg.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge