Markus Lundgren
Modeling Disease Progression Trajectories from Longitudinal Observational Data
Dec 09, 2020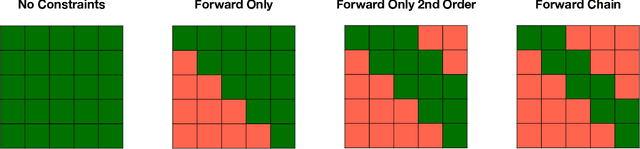
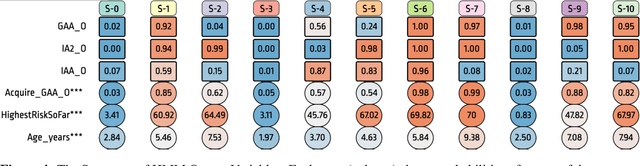
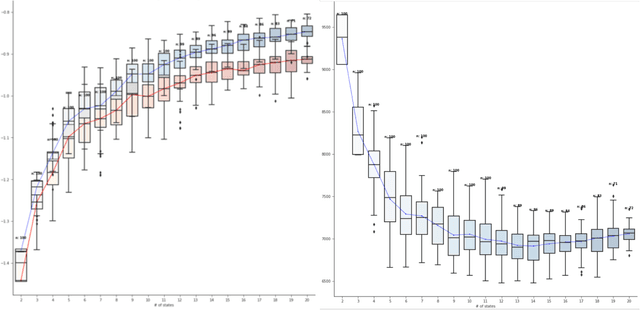
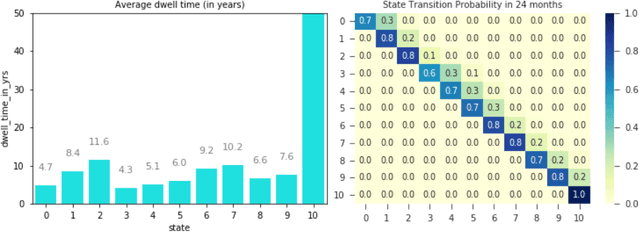

Abstract:Analyzing disease progression patterns can provide useful insights into the disease processes of many chronic conditions. These analyses may help inform recruitment for prevention trials or the development and personalization of treatments for those affected. We learn disease progression patterns using Hidden Markov Models (HMM) and distill them into distinct trajectories using visualization methods. We apply it to the domain of Type 1 Diabetes (T1D) using large longitudinal observational data from the T1DI study group. Our method discovers distinct disease progression trajectories that corroborate with recently published findings. In this paper, we describe the iterative process of developing the model. These methods may also be applied to other chronic conditions that evolve over time.
DPVis: Visual Exploration of Disease Progression Pathways
Apr 26, 2019Abstract:Clinical researchers use disease progression modeling algorithms to predict future patient status and characterize progression patterns. One approach for disease progression modeling is to describe patient status using a small number of states that represent distinctive distributions over a set of observed measures. Hidden Markov models (HMMs) and its variants are a class of models that both discover these states and make predictions concerning future states for new patients. HMMs can be trained using longitudinal observations of subjects from large-scale cohort studies, clinical trials, and electronic health records. Despite the advantages of using the algorithms for discovering interesting patterns, it still remains challenging for medical experts to interpret model outputs, complex modeling parameters, and clinically make sense of the patterns. To tackle this problem, we conducted a design study with physician scientists, statisticians, and visualization experts, with the goal to investigate disease progression pathways of certain chronic diseases, namely type 1 diabetes (T1D), Huntington's disease, Parkinson's disease, and chronic obstructive pulmonary disease (COPD). As a result, we introduce DPVis which seamlessly integrates model parameters and outcomes of HMMs into interpretable, and interactive visualizations. In this study, we demonstrate that DPVis is successful in evaluating disease progression models, visually summarizing disease states, interactively exploring disease progression patterns, and designing and comparing clinically relevant subgroup cohorts by introducing a case study on observation data from clinical studies of T1D.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge