Manan Lalit
Unsupervised Learning of Object-Centric Embeddings for Cell Instance Segmentation in Microscopy Images
Oct 12, 2023
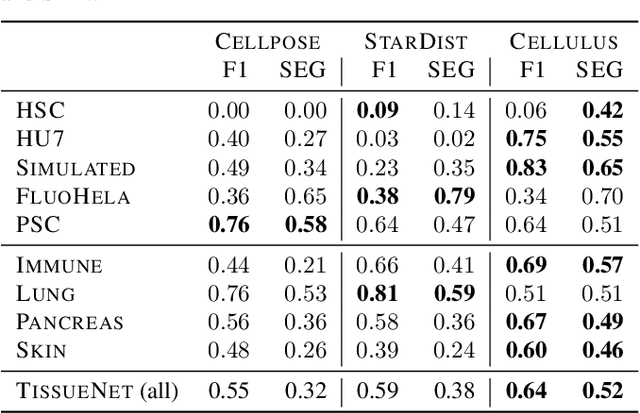
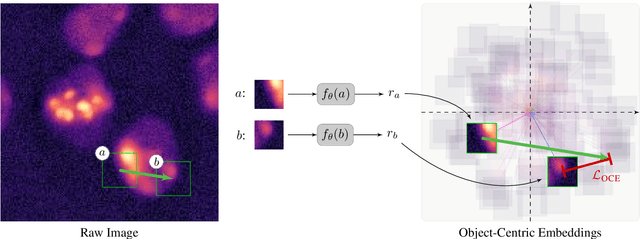
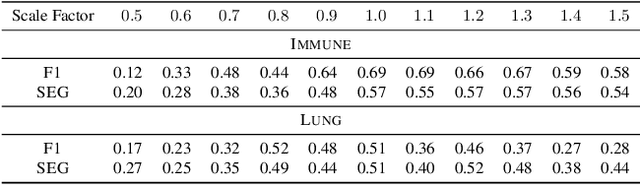
Abstract:Segmentation of objects in microscopy images is required for many biomedical applications. We introduce object-centric embeddings (OCEs), which embed image patches such that the spatial offsets between patches cropped from the same object are preserved. Those learnt embeddings can be used to delineate individual objects and thus obtain instance segmentations. Here, we show theoretically that, under assumptions commonly found in microscopy images, OCEs can be learnt through a self-supervised task that predicts the spatial offset between image patches. Together, this forms an unsupervised cell instance segmentation method which we evaluate on nine diverse large-scale microscopy datasets. Segmentations obtained with our method lead to substantially improved results, compared to state-of-the-art baselines on six out of nine datasets, and perform on par on the remaining three datasets. If ground-truth annotations are available, our method serves as an excellent starting point for supervised training, reducing the required amount of ground-truth needed by one order of magnitude, thus substantially increasing the practical applicability of our method. Source code is available at https://github.com/funkelab/cellulus.
Embedding-based Instance Segmentation of Microscopy Images
Jan 25, 2021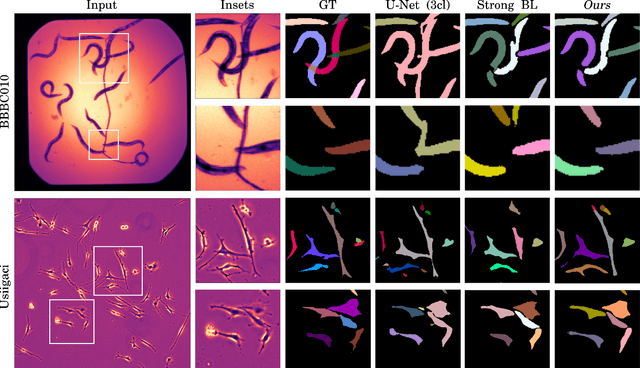

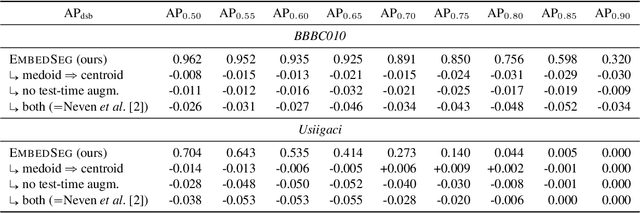
Abstract:Automatic detection and segmentation of objects in microscopy images is important for many biological applications. In the domain of natural images, and in particular in the context of city street scenes, embedding-based instance segmentation leads to high-quality results. Inspired by this line of work, we introduce EmbedSeg, an end-to-end trainable deep learning method based on the work by Neven et al. While their approach embeds each pixel to the centroid of any given instance, in EmbedSeg, motivated by the complex shapes of biological objects, we propose to use the medoid instead. Additionally, we make use of a test-time augmentation scheme, and show that both suggested modifications improve the instance segmentation performance on biological microscopy datasets notably. We demonstrate that embedding-based instance segmentation achieves competitive results in comparison to state-of-the-art methods on diverse and biologically relevant microscopy datasets. Finally, we show that the overall pipeline has a small enough memory footprint to be used on virtually all CUDA enabled laptop hardware. Our open-source implementation is available at github.com/juglab/EmbedSeg.
Leveraging Self-supervised Denoising for Image Segmentation
Dec 13, 2019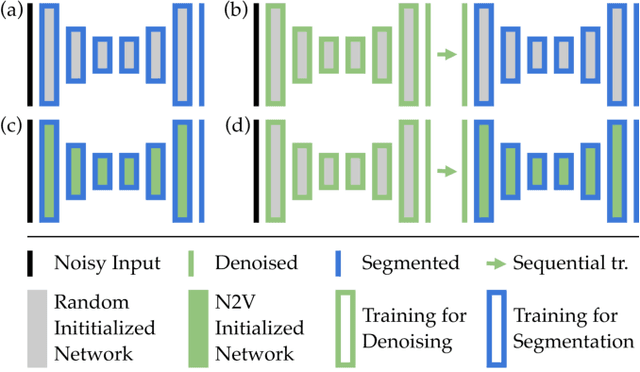
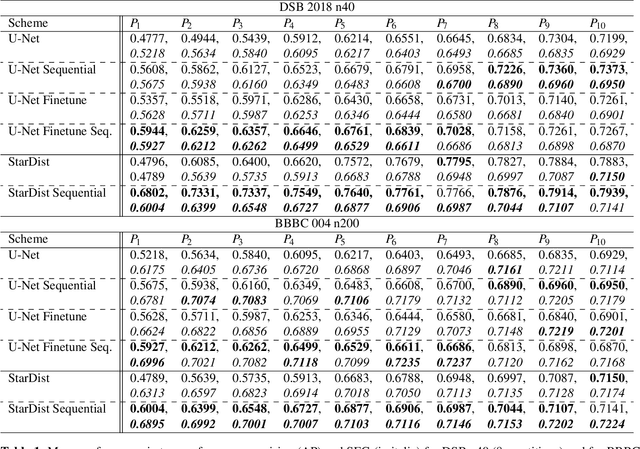
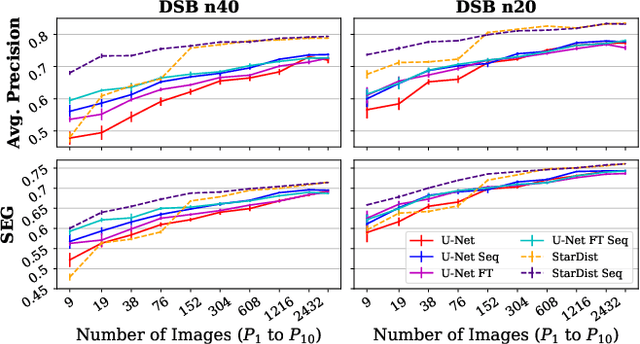
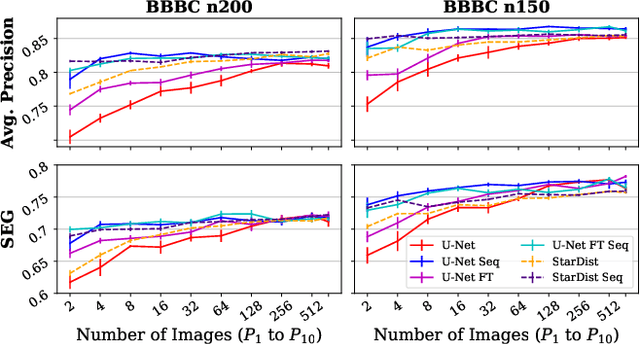
Abstract:Deep learning (DL) has arguably emerged as the method of choice for the detection and segmentation of biological structures in microscopy images. However, DL typically needs copious amounts of annotated training data that is for biomedical projects typically not available and excessively expensive to generate. Additionally, tasks become harder in the presence of noise, requiring even more high-quality training data. Hence, we propose to use denoising networks to improve the performance of other DL-based image segmentation methods. More specifically, we present ideas on how state-of-the-art self-supervised CARE networks can improve cell/nuclei segmentation in microscopy data. Using two state-of-the-art baseline methods, U-Net and StarDist, we show that our ideas consistently improve the quality of resulting segmentations, especially when only limited training data for noisy micrographs are available.
Fully Unsupervised Probabilistic Noise2Void
Nov 27, 2019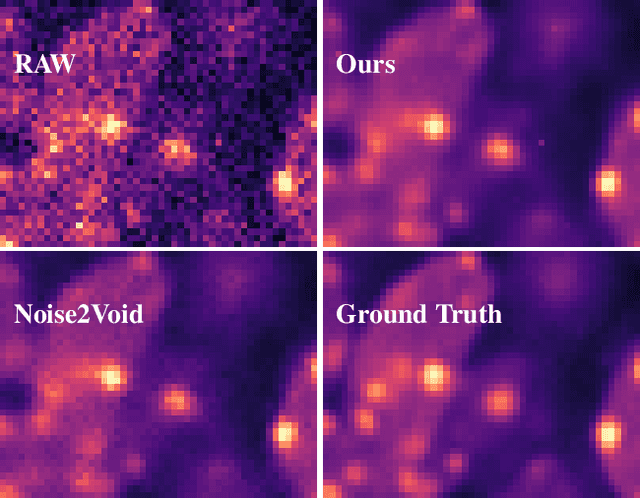



Abstract:Image denoising is the first step in many biomedical image analysis pipelines and Deep Learning (DL) based methods are currently best performing. A new category of DL methods such as Noise2Void or Noise2Self can be used fully unsupervised, requiring nothing but the noisy data. However, this comes at the price of reduced reconstruction quality. The recently proposed Probabilistic Noise2Void (PN2V) improves results, but requires an additional noise model for which calibration data needs to be acquired. Here, we present improvements to PN2V that (i) replace histogram based noise models by parametric noise models, and (ii) show how suitable noise models can be created even in the absence of calibration data. This is a major step since it actually renders PN2V fully unsupervised. We demonstrate that all proposed improvements are not only academic but indeed relevant.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge