Laura L. Elo
RPA: Probabilistic analysis of probe performance and robust summarization
Apr 06, 2013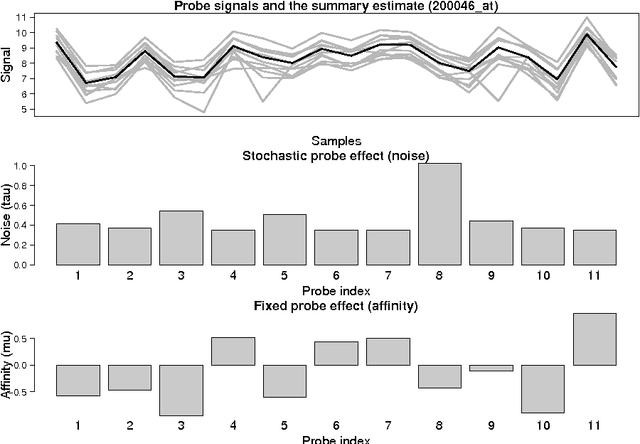
Abstract:Probe-level models have led to improved performance in microarray studies but the various sources of probe-level contamination are still poorly understood. Data-driven analysis of probe performance can be used to quantify the uncertainty in individual probes and to highlight the relative contribution of different noise sources. Improved understanding of the probe-level effects can lead to improved preprocessing techniques and microarray design. We have implemented probabilistic tools for probe performance analysis and summarization on short oligonucleotide arrays. In contrast to standard preprocessing approaches, the methods provide quantitative estimates of probe-specific noise and affinity terms and tools to investigate these parameters. Tools to incorporate prior information of the probes in the analysis are provided as well. Comparisons to known probe-level error sources and spike-in data sets validate the approach. Implementation is freely available in R/BioConductor: http://www.bioconductor.org/packages/release/bioc/html/RPA.html
Fully scalable online-preprocessing algorithm for short oligonucleotide microarray atlases
Dec 27, 2012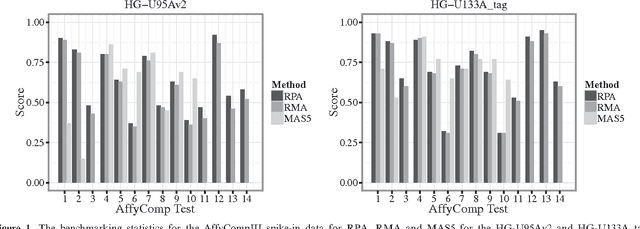
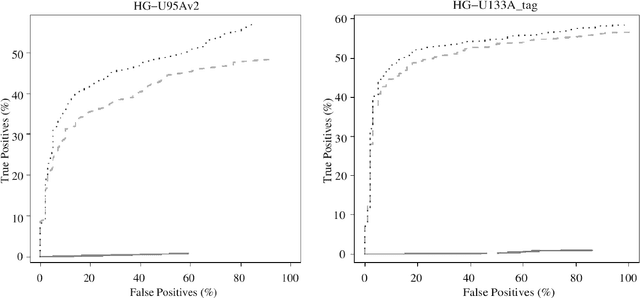
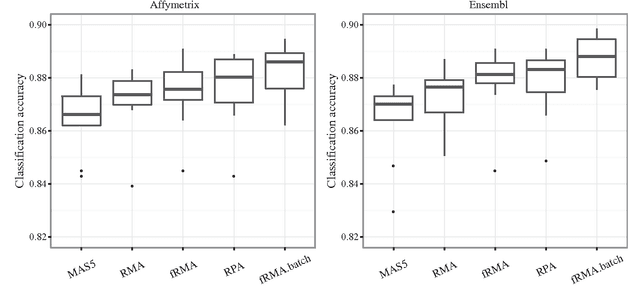
Abstract:Accumulation of standardized data collections is opening up novel opportunities for holistic characterization of genome function. The limited scalability of current preprocessing techniques has, however, formed a bottleneck for full utilization of contemporary microarray collections. While short oligonucleotide arrays constitute a major source of genome-wide profiling data, scalable probe-level preprocessing algorithms have been available only for few measurement platforms based on pre-calculated model parameters from restricted reference training sets. To overcome these key limitations, we introduce a fully scalable online-learning algorithm that provides tools to process large microarray atlases including tens of thousands of arrays. Unlike the alternatives, the proposed algorithm scales up in linear time with respect to sample size and is readily applicable to all short oligonucleotide platforms. This is the only available preprocessing algorithm that can learn probe-level parameters based on sequential hyperparameter updates at small, consecutive batches of data, thus circumventing the extensive memory requirements of the standard approaches and opening up novel opportunities to take full advantage of contemporary microarray data collections. Moreover, using the most comprehensive data collections to estimate probe-level effects can assist in pinpointing individual probes affected by various biases and provide new tools to guide array design and quality control. The implementation is freely available in R/Bioconductor at http://www.bioconductor.org/packages/devel/bioc/html/RPA.html
* 20 pages, 3 figures, 1 supplementary PDF
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge