Krishna R Narayanan
BayReL: Bayesian Relational Learning for Multi-omics Data Integration
Oct 22, 2020
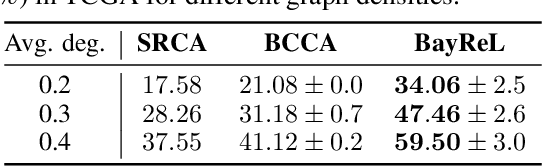
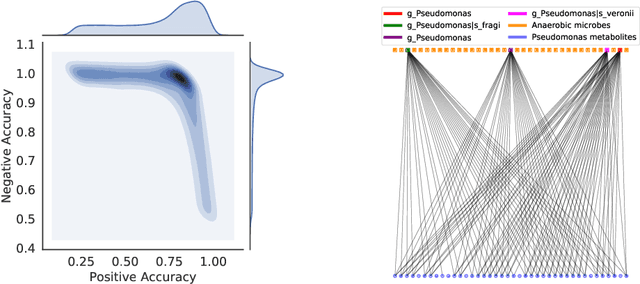
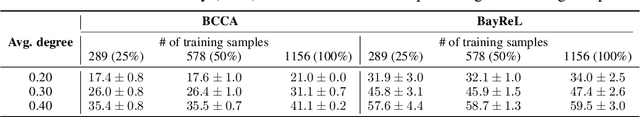
Abstract:High-throughput molecular profiling technologies have produced high-dimensional multi-omics data, enabling systematic understanding of living systems at the genome scale. Studying molecular interactions across different data types helps reveal signal transduction mechanisms across different classes of molecules. In this paper, we develop a novel Bayesian representation learning method that infers the relational interactions across multi-omics data types. Our method, Bayesian Relational Learning (BayReL) for multi-omics data integration, takes advantage of a priori known relationships among the same class of molecules, modeled as a graph at each corresponding view, to learn view-specific latent variables as well as a multi-partite graph that encodes the interactions across views. Our experiments on several real-world datasets demonstrate enhanced performance of BayReL in inferring meaningful interactions compared to existing baselines.
Variational Graph Recurrent Neural Networks
Sep 09, 2019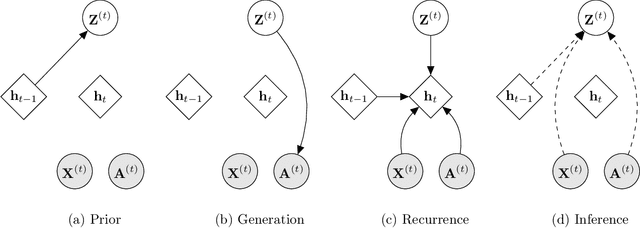
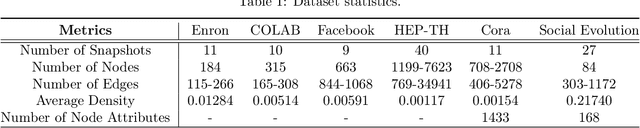
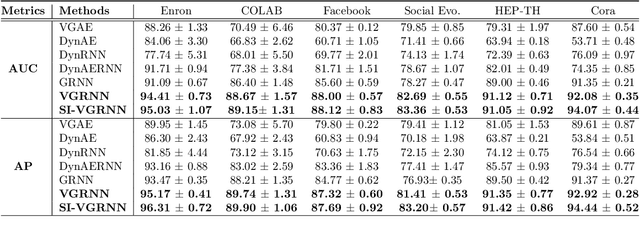
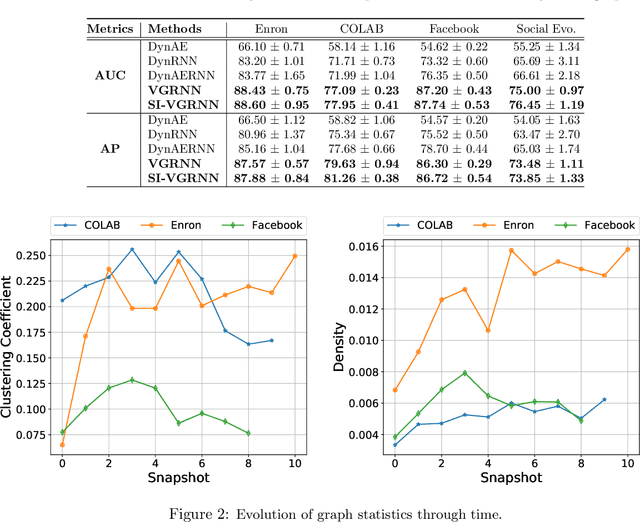
Abstract:Representation learning over graph structured data has been mostly studied in static graph settings while efforts for modeling dynamic graphs are still scant. In this paper, we develop a novel hierarchical variational model that introduces additional latent random variables to jointly model the hidden states of a graph recurrent neural network (GRNN) to capture both topology and node attribute changes in dynamic graphs. We argue that the use of high-level latent random variables in this variational GRNN (VGRNN) can better capture potential variability observed in dynamic graphs as well as the uncertainty of node latent representation. With semi-implicit variational inference developed for this new VGRNN architecture (SI-VGRNN), we show that flexible non-Gaussian latent representations can further help dynamic graph analytic tasks. Our experiments with multiple real-world dynamic graph datasets demonstrate that SI-VGRNN and VGRNN consistently outperform the existing baseline and state-of-the-art methods by a significant margin in dynamic link prediction.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge