Kaifeng Pang
Cross-Slice Attention and Evidential Critical Loss for Uncertainty-Aware Prostate Cancer Detection
Jul 01, 2024

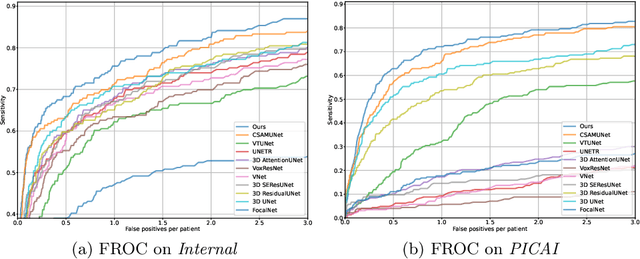
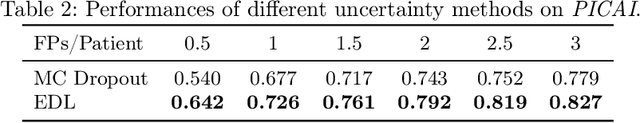
Abstract:Current deep learning-based models typically analyze medical images in either 2D or 3D albeit disregarding volumetric information or suffering sub-optimal performance due to the anisotropic resolution of MR data. Furthermore, providing an accurate uncertainty estimation is beneficial to clinicians, as it indicates how confident a model is about its prediction. We propose a novel 2.5D cross-slice attention model that utilizes both global and local information, along with an evidential critical loss, to perform evidential deep learning for the detection in MR images of prostate cancer, one of the most common cancers and a leading cause of cancer-related death in men. We perform extensive experiments with our model on two different datasets and achieve state-of-the-art performance in prostate cancer detection along with improved epistemic uncertainty estimation. The implementation of the model is available at https://github.com/aL3x-O-o-Hung/GLCSA_ECLoss.
CSAM: A 2.5D Cross-Slice Attention Module for Anisotropic Volumetric Medical Image Segmentation
Nov 08, 2023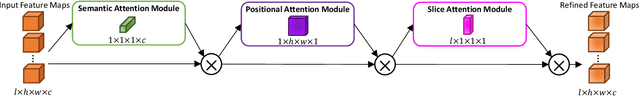
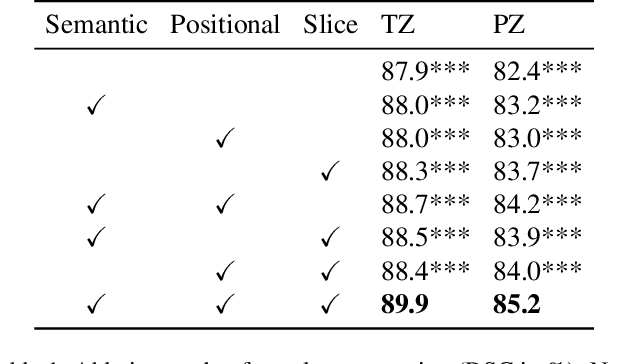
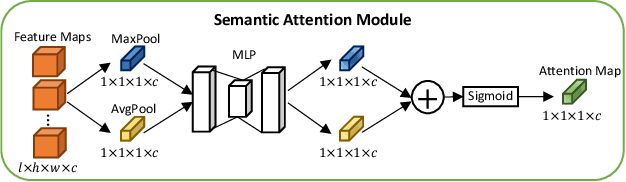
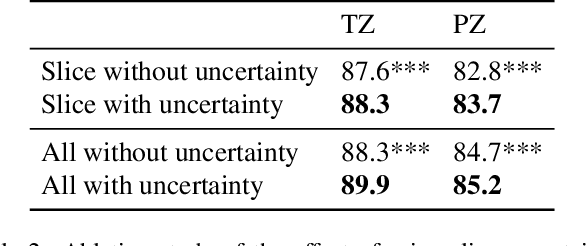
Abstract:A large portion of volumetric medical data, especially magnetic resonance imaging (MRI) data, is anisotropic, as the through-plane resolution is typically much lower than the in-plane resolution. Both 3D and purely 2D deep learning-based segmentation methods are deficient in dealing with such volumetric data since the performance of 3D methods suffers when confronting anisotropic data, and 2D methods disregard crucial volumetric information. Insufficient work has been done on 2.5D methods, in which 2D convolution is mainly used in concert with volumetric information. These models focus on learning the relationship across slices, but typically have many parameters to train. We offer a Cross-Slice Attention Module (CSAM) with minimal trainable parameters, which captures information across all the slices in the volume by applying semantic, positional, and slice attention on deep feature maps at different scales. Our extensive experiments using different network architectures and tasks demonstrate the usefulness and generalizability of CSAM. Associated code is available at https://github.com/aL3x-O-o-Hung/CSAM.
PartDiff: Image Super-resolution with Partial Diffusion Models
Jul 21, 2023Abstract:Denoising diffusion probabilistic models (DDPMs) have achieved impressive performance on various image generation tasks, including image super-resolution. By learning to reverse the process of gradually diffusing the data distribution into Gaussian noise, DDPMs generate new data by iteratively denoising from random noise. Despite their impressive performance, diffusion-based generative models suffer from high computational costs due to the large number of denoising steps.In this paper, we first observed that the intermediate latent states gradually converge and become indistinguishable when diffusing a pair of low- and high-resolution images. This observation inspired us to propose the Partial Diffusion Model (PartDiff), which diffuses the image to an intermediate latent state instead of pure random noise, where the intermediate latent state is approximated by the latent of diffusing the low-resolution image. During generation, Partial Diffusion Models start denoising from the intermediate distribution and perform only a part of the denoising steps. Additionally, to mitigate the error caused by the approximation, we introduce "latent alignment", which aligns the latent between low- and high-resolution images during training. Experiments on both magnetic resonance imaging (MRI) and natural images show that, compared to plain diffusion-based super-resolution methods, Partial Diffusion Models significantly reduce the number of denoising steps without sacrificing the quality of generation.
MAg: a simple learning-based patient-level aggregation method for detecting microsatellite instability from whole-slide images
Jan 13, 2022

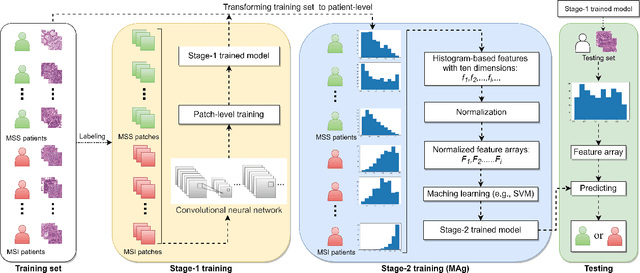

Abstract:The prediction of microsatellite instability (MSI) and microsatellite stability (MSS) is essential in predicting both the treatment response and prognosis of gastrointestinal cancer. In clinical practice, a universal MSI testing is recommended, but the accessibility of such a test is limited. Thus, a more cost-efficient and broadly accessible tool is desired to cover the traditionally untested patients. In the past few years, deep-learning-based algorithms have been proposed to predict MSI directly from haematoxylin and eosin (H&E)-stained whole-slide images (WSIs). Such algorithms can be summarized as (1) patch-level MSI/MSS prediction, and (2) patient-level aggregation. Compared with the advanced deep learning approaches that have been employed for the first stage, only the na\"ive first-order statistics (e.g., averaging and counting) were employed in the second stage. In this paper, we propose a simple yet broadly generalizable patient-level MSI aggregation (MAg) method to effectively integrate the precious patch-level information. Briefly, the entire probabilistic distribution in the first stage is modeled as histogram-based features to be fused as the final outcome with machine learning (e.g., SVM). The proposed MAg method can be easily used in a plug-and-play manner, which has been evaluated upon five broadly used deep neural networks: ResNet, MobileNetV2, EfficientNet, Dpn and ResNext. From the results, the proposed MAg method consistently improves the accuracy of patient-level aggregation for two publicly available datasets. It is our hope that the proposed method could potentially leverage the low-cost H&E based MSI detection method. The code of our work has been made publicly available at https://github.com/Calvin-Pang/MAg.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge