Jun Itami
Unsupervised Brain Abnormality Detection Using High Fidelity Image Reconstruction Networks
Jun 02, 2020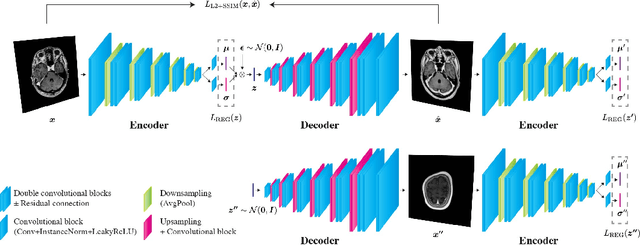
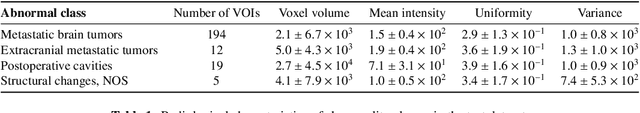

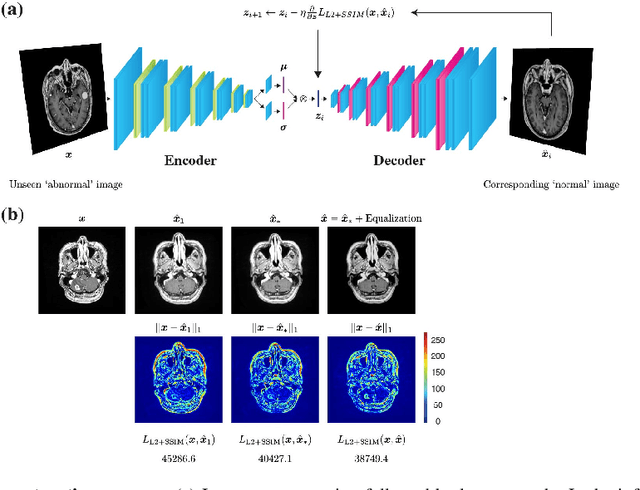
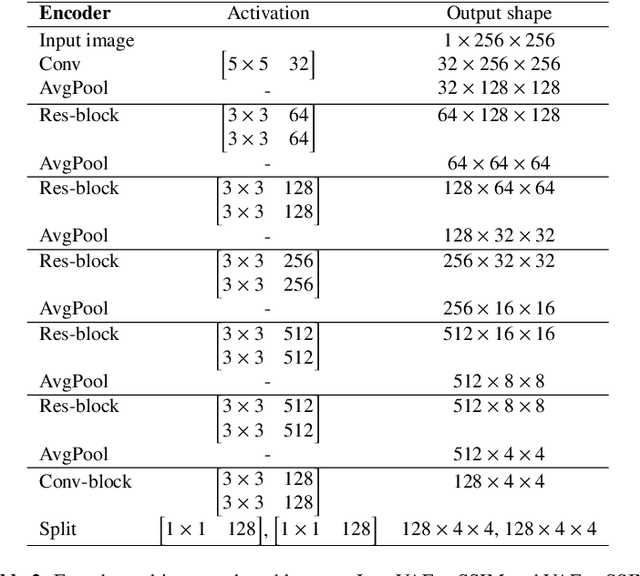
Abstract:Recent advances in deep learning have facilitated near-expert medical image analysis. Supervised learning is the mainstay of current approaches, though its success requires the use of large, fully labeled datasets. However, in real-world medical practice, previously unseen disease phenotypes are encountered that have not been defined a priori in finite-size datasets. Unsupervised learning, a hypothesis-free learning framework, may play a complementary role to supervised learning. Here, we demonstrate a novel framework for voxel-wise abnormality detection in brain magnetic resonance imaging (MRI), which exploits an image reconstruction network based on an introspective variational autoencoder trained with a structural similarity constraint. The proposed network learns a latent representation for "normal" anatomical variation using a series of images that do not include annotated abnormalities. After training, the network can map unseen query images to positions in the latent space, and latent variables sampled from those positions can be mapped back to the image space to yield normal-looking replicas of the input images. Finally, the network considers abnormality scores, which are designed to reflect differences at several image feature levels, in order to locate image regions that may contain abnormalities. The proposed method is evaluated on a comprehensively annotated dataset spanning clinically significant structural abnormalities of the brain parenchyma in a population having undergone radiotherapy for brain metastasis, demonstrating that it is particularly effective for contrast-enhanced lesions, i.e., metastatic brain tumors and extracranial metastatic tumors.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge