Juan Yang
Structure-to-Image: Zero-Shot Depth Estimation in Colonoscopy via High-Fidelity Sim-to-Real Adaptation
Feb 25, 2026Abstract:Monocular depth estimation (MDE) for colonoscopy is hampered by the domain gap between simulated and real-world images. Existing image-to-image translation methods, which use depth as a posterior constraint, often produce structural distortions and specular highlights by failing to balance realism with structure consistency. To address this, we propose a Structure-to-Image paradigm that transforms the depth map from a passive constraint into an active generative foundation. We are the first to introduce phase congruency to colonoscopic domain adaptation and design a cross-level structure constraint to co-optimize geometric structures and fine-grained details like vascular textures. In zero-shot evaluations conducted on a publicly available phantom dataset, the MDE model that was fine-tuned on our generated data achieved a maximum reduction of 44.18% in RMSE compared to competing methods. Our code is available at https://github.com/YyangJJuan/PC-S2I.git.
LKCell: Efficient Cell Nuclei Instance Segmentation with Large Convolution Kernels
Jul 25, 2024


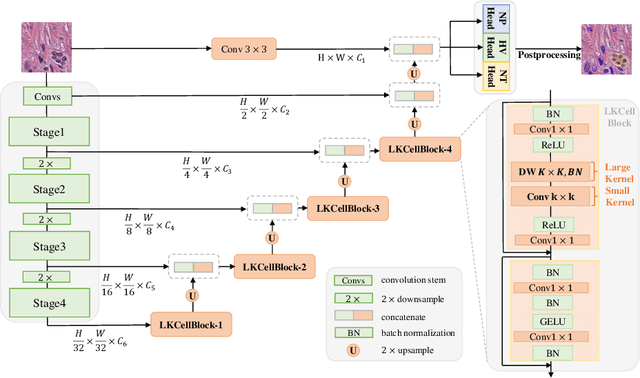
Abstract:The segmentation of cell nuclei in tissue images stained with the blood dye hematoxylin and eosin (H$\&$E) is essential for various clinical applications and analyses. Due to the complex characteristics of cellular morphology, a large receptive field is considered crucial for generating high-quality segmentation. However, previous methods face challenges in achieving a balance between the receptive field and computational burden. To address this issue, we propose LKCell, a high-accuracy and efficient cell segmentation method. Its core insight lies in unleashing the potential of large convolution kernels to achieve computationally efficient large receptive fields. Specifically, (1) We transfer pre-trained large convolution kernel models to the medical domain for the first time, demonstrating their effectiveness in cell segmentation. (2) We analyze the redundancy of previous methods and design a new segmentation decoder based on large convolution kernels. It achieves higher performance while significantly reducing the number of parameters. We evaluate our method on the most challenging benchmark and achieve state-of-the-art results (0.5080 mPQ) in cell nuclei instance segmentation with only 21.6% FLOPs compared with the previous leading method. Our source code and models are available at https://github.com/hustvl/LKCell.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge